Hello,
I’m trying to fit the side band region of a 2D data set. The region of the data distribution is at x[-10,10] and y[-10,10]. The side band is defined to be -10<x<10 and -10<y<0 using
x.setRange("SB1",-10,+10) ;
y.setRange("SB1",-10,0) ;
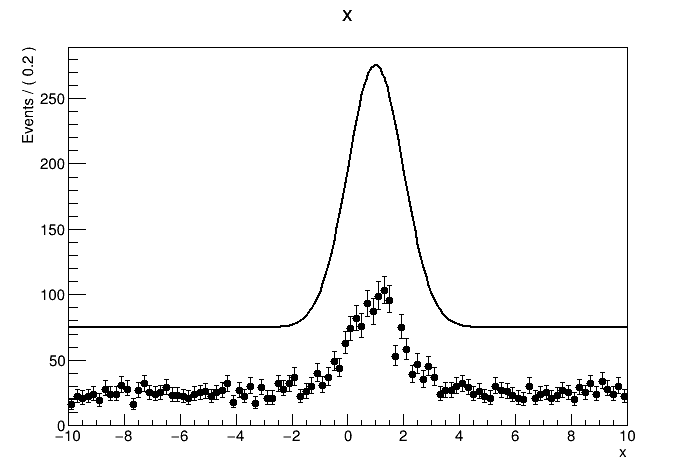
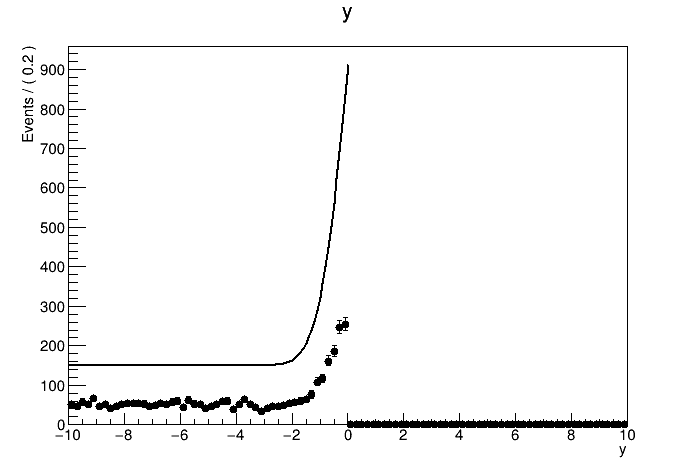
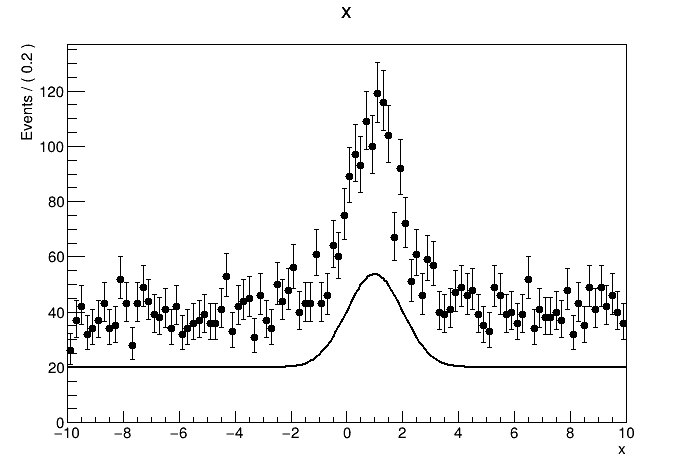
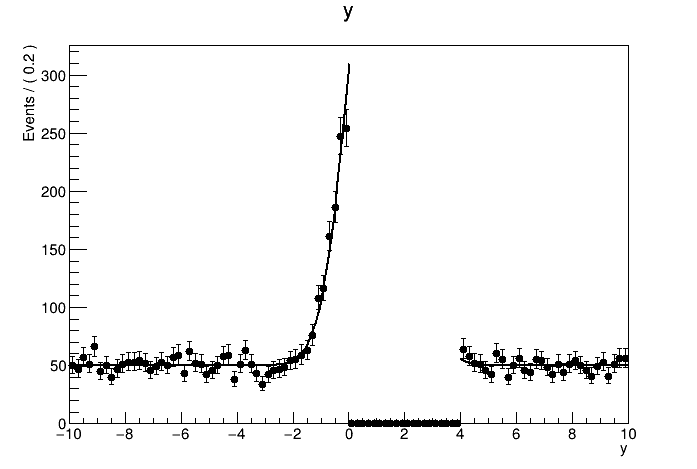
The result of the fit is correct. But when I tried to plot out the data with the pdf model withNormRange("SB1"), the normalization of the model seems to be totally wrong. How should I plot the result of a side band fit with the correct normalization?
See the attachment:
using namespace RooFit ;
void sideBandEx()
{
// C r e a t e 2 D p d f a n d d a t a
// -------------------------------------------
// Define observables x,y
RooRealVar x("x","x",-10,10) ;
RooRealVar y("y","y",-10,10) ;
// Construct the signal pdf gauss(x)*gauss(y)
RooRealVar mx("mx","mx",1,-10,10) ;
RooRealVar my("my","my",1,-10,10) ;
RooGaussian gx("gx","gx",x,mx,RooConst(1)) ;
RooGaussian gy("gy","gy",y,my,RooConst(1)) ;
RooProdPdf sig("sig","sig",gx,gy) ;
// Construct the background pdf (flat in x,y)
RooPolynomial px("px","px",x) ;
RooPolynomial py("py","py",y) ;
RooProdPdf bkg("bkg","bkg",px,py) ;
// Construct the composite model sig+bkg
RooRealVar f("f","f",0.,1.) ;
RooAddPdf model("model","model",RooArgList(sig,bkg),f) ;
// Sample 10000 events in (x,y) from the model
RooDataSet* modelData = model.generate(RooArgSet(x,y),10000) ;
x.setRange("SB1",-10,+10) ;
y.setRange("SB1",-10,0) ;
RooFitResult* r_sb12 = model.fitTo(*modelData,Range("SB1"),Save()) ;
// Plot the results
RooPlot* frame01 = x.frame(Title("x"));
modelData->plotOn(frame01,CutRange("SB1"));
model.plotOn(frame01,LineColor(1),LineWidth(2),ProjectionRange("SB1"),NormRange("SB1"));
RooPlot* frame02 = y.frame(Title("y"));
modelData->plotOn(frame02,CutRange("SB1"));
model.plotOn(frame02,LineColor(1),LineWidth(2),ProjectionRange("SB1"),NormRange("SB1"));
TCanvas *c1 = new TCanvas("c1","",1);
frame01->Draw();
TCanvas *c2 = new TCanvas("c2","",1);
frame02->Draw();
r_sb12->Print() ;
}
The plots:
The output messages:
[#1] INFO:Minization -- RooMinimizer::optimizeConst: deactivating const optimization
[#1] INFO:Plotting -- RooTreeData::plotOn: plotting 3335 events out of 10000 total events
[#1] INFO:Plotting -- RooAbsPdf::plotOn(model) p.d.f was fitted in range and no explicit plot range was specified, using fit range as default
[#1] INFO:Plotting -- RooAbsPdf::plotOn(model) only plotting range 'fit_nll_model_modelData,fit_nll_model_modelData'
[#1] INFO:Plotting -- RooAbsPdf::plotOn(model) p.d.f. curve is normalized using explicit choice of ranges 'SB1'
[#0] WARNING:Plotting -- RooHist::getFitRangeNEvt() WARNING: Number of normalization events associated to histogram is not equal to number of events in histogram
due cut made in RooAbsData::plotOn() call. Automatic normalization over sub-range of plot variable assumes
that the effect of that cut is uniform across the plot, which may be an incorrect assumption. To be sure of
correct normalization explicit pass normalization information to RooAbsPdf::plotOn() call using Normalization()
[#1] INFO:Plotting -- RooAbsReal::plotOn(model) plot on x integrates over variables (y) in range SB1
[#0] ERROR:InputArguments -- RooArgSet::checkForDup: ERROR argument with name model_Int[x,y|fit_nll_model_modelData]_Norm[x,y] is already in this set
[#1] INFO:Plotting -- RooAbsReal::plotOn(model) plot on x integrates over variables (y) in range SB1
[#0] ERROR:InputArguments -- RooArgSet::checkForDup: ERROR argument with name model_Int[x,y|fit_nll_model_modelData]_Norm[x,y] is already in this set
[#1] INFO:Plotting -- RooTreeData::plotOn: plotting 3335 events out of 10000 total events
[#1] INFO:Plotting -- RooAbsPdf::plotOn(model) p.d.f was fitted in range and no explicit plot range was specified, using fit range as default
[#1] INFO:Plotting -- RooAbsPdf::plotOn(model) only plotting range 'fit_nll_model_modelData,fit_nll_model_modelData'
[#1] INFO:Plotting -- RooAbsPdf::plotOn(model) p.d.f. curve is normalized using explicit choice of ranges 'SB1'
[#0] WARNING:Plotting -- RooHist::getFitRangeNEvt() WARNING: Number of normalization events associated to histogram is not equal to number of events in histogram
due cut made in RooAbsData::plotOn() call. Automatic normalization over sub-range of plot variable assumes
that the effect of that cut is uniform across the plot, which may be an incorrect assumption. To be sure of
correct normalization explicit pass normalization information to RooAbsPdf::plotOn() call using Normalization()
[#1] INFO:Plotting -- RooAbsReal::plotOn(model) plot on y integrates over variables (x) in range SB1
[#0] ERROR:InputArguments -- RooArgSet::checkForDup: ERROR argument with name model_Int[x,y|fit_nll_model_modelData]_Norm[x,y] is already in this set
[#1] INFO:Plotting -- RooAbsReal::plotOn(model) plot on y integrates over variables (x) in range SB1
[#0] ERROR:InputArguments -- RooArgSet::checkForDup: ERROR argument with name model_Int[x,y|fit_nll_model_modelData]_Norm[x,y] is already in this set