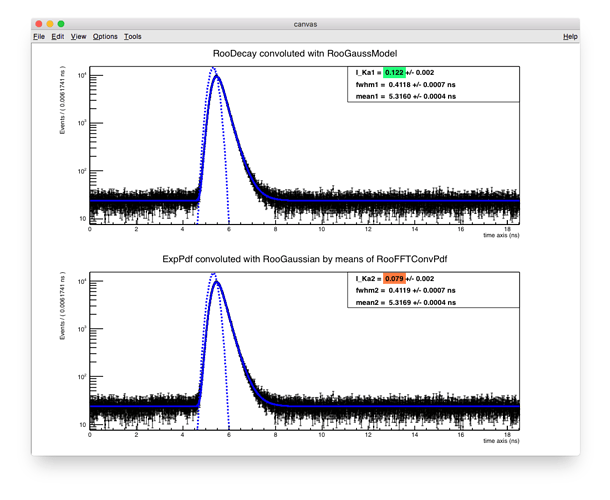
Generally speaking in RooFit the convolution operation of a superposition of exponential decays with a Gaussian function can be carried out in two different ways. Either we are using analytical convolution (RooDecay and RooGaussModel) or convolution of a custom decay derived from RooAbsPdf with one of numeric convolution operator classes – RooFFTConvPdf or RooNumConvPdf.
We tried implementing both techniques in our software and compared fit results with the existing products on the market (we do positron annihilation lifetime fitting). It turns out that RooDecay’s convoluted with RooGaussModel provide correct parameter values, whereas custom RooAbsPdf decays convoluted by means of RooFFTConvPdf give one incorrect parameter value.
A compact ROOT macro that illustrates the problem is attached to this topic. Here we import a histogram from file and fit it with a sum of two exponential decays convoluted with a single gauss (in the end also adding constant background). Two above mentioned approaches are implemented in the macro. Fit clearly demonstrates that the intensity of second exponential component I_Ka is incorrect (7.9%) for RooFFTConvPdf case. Correct value of 12% is given by RooDecay and RooGaussModel. We double checked this by performing the fit of the same sectrum with third-party positron annihilation lifetime fitting software (LT10 and PALSfit).
I’ve played with different binning and buffer settings of RooFFTConvPdf and tried to use RooNumConvPdf instead. No luck. Fit itself converges and looks good. But it can be a buggy calculation of these RooAddPdf components’ intensity values.
Clarification of this issue is extremely important for us. Looking forward to hearing back from you. Code tested with ROOT 6.11/02 and RooFit v3.60.
convtest.zip (10.4 KB)