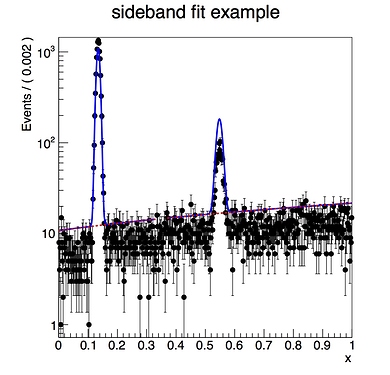
Dear experts,
I am running into the following problem related to fitting in sub-ranges (sidebands). I created a minimal example macro that reproduces my problem. The rootfile and macro are attached.
setup
I have some spectrum where there are two Gaussian peaks and some smooth polynomial background in between them (and underneath both peaks). I want to fit the spectrum but with some section in the middle region excluded. The idea is to use the sidebands to completely constrain the shape and normalization over the excluded region given this model.
problem
The fit runs perfectly fine if I fit on the full range:
model.fitTo(*data);
However, it fails if I run the fit on left+right sidebands:
model.fitTo(*data,Range("SBLeft,SBRight"));
The core parts regarding the model definition and fitting looks like the following:
RooRealVar x("x","x",0.,1.);
RooRealVar mean_1("mean_1","mean 1",0.135,0.134,0.136);
RooRealVar sigma_1("sigma_1","sigma 1",0.006,0.005,0.007);
RooGaussian gaus_1("gaus_1","gaus 1",x,mean_1,sigma_1);
RooRealVar mean_2("mean_2","mean 2",0.548,0.547,0.549);
RooRealVar sigma_2("sigma_2","sigma 2",0.0098,0.0097,0.0099);
RooGaussian gaus_2("gaus_2","gaus 2",x,mean_2,sigma_2);
RooRealVar a0("a0","a0",1.,0.,2.);
RooPolynomial bkg("bkg","bkg",x,RooArgSet(a0));
RooRealVar ngaus_1("ngaus_1","ngaus 1",8000.,100,100000);
RooRealVar ngaus_2("ngaus_2","ngaus ",2000.,100,100000);
RooRealVar nbkg("nbkg","nbkg",8000.,10,10000);
RooAddPdf model("model","model",RooArgList(gaus_1,gaus_2,bkg),RooArgList(ngaus_1,ngaus_2,nbkg));
x.setRange("SBLeft", 0., 0.33);
x.setRange("SBRight", 0.37 , 1.);
//RooFitResult* result = model.fitTo(*data); // success
RooFitResult* result = model.fitTo(*data,Range("SBLeft,SBRight")); // fail
my guess of the cause
After reading some posts and playing with it, I think the cause of the problem is the following:
When fitting multiple ranges, pdfs are normalized and log-likelihoods are calculated over each individual range before they are summed and fed to the minimizer. This will be a problem when normalizing over, e.g. SBLeft, for the component gaus_2 since it is so far in the tail, the normalization will be 0 in this sub-range, creating a problem.
questions
Hence, my questions are:
- Is my interpretation above of why it fails correct?
- What is the proper way to handle this situation with RooFit?
Thank you very much!
Regards,
Yunjie
minimal_example_fit.C (1.9 KB)
minimal_output_generated.root (123.6 KB)
P.S. I am working with ROOT 6.12.04 and RooFit v3.60