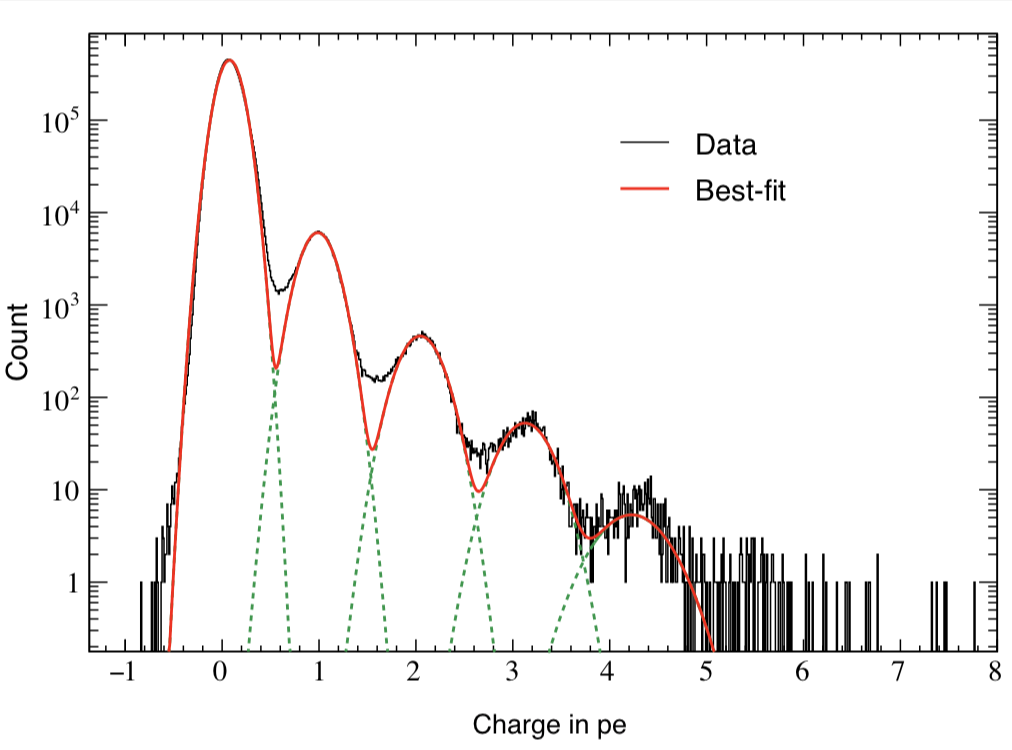
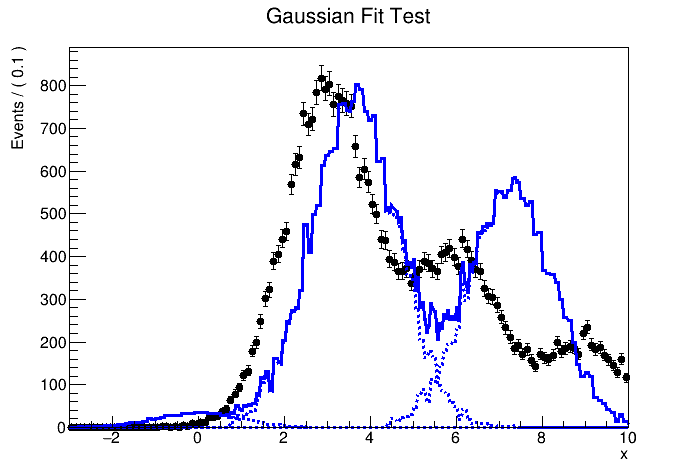
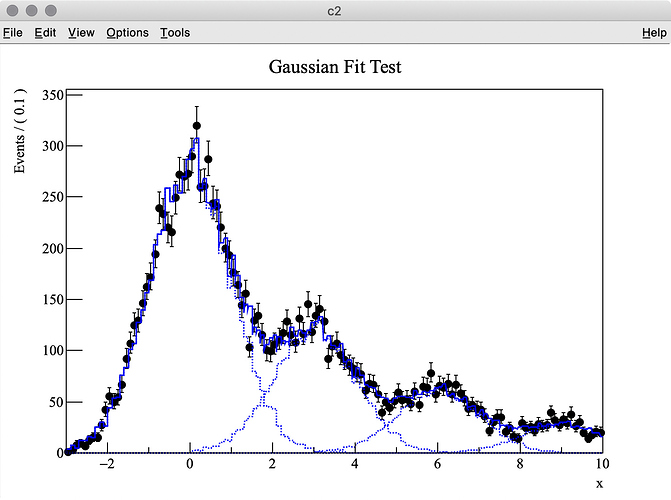
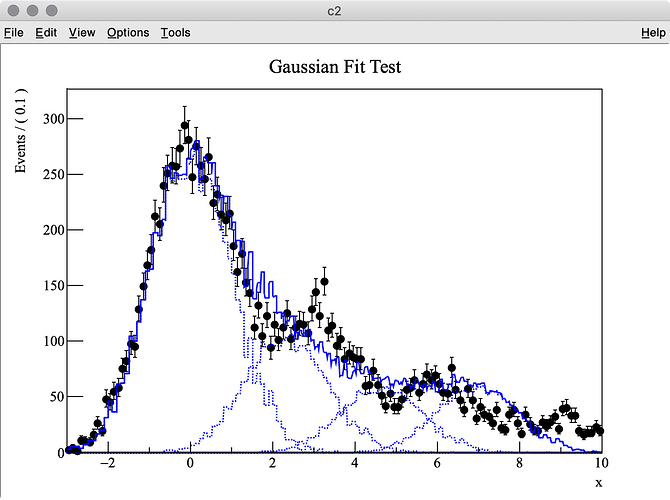
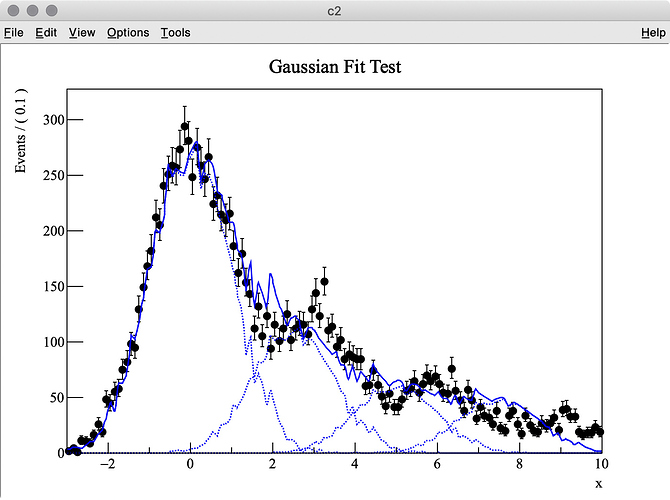
I try to fit simulation data (TH1 “h2”) containing three Gaussian peaks with a model probability distribution function (PDF) consisting of a sum of three Gaussian PDFs. The first Gaussian PDF is located at x = 0 (TH1 “h1”), the other two PDF shapes are identical with the first one, but they are horizontally shifted by dx and 2 * dx by using RooFormulaVar.
When making the model PDF, I set four parameters below :
dx : the x offset of shifted PDFs (2 * dx to be used for the second one)
p0, p1, p2 : scaling parameters for individual PDFs
pdf2.C (3.4 KB)
$ root
root [0] .x pdf2.C(1000000, 10000, 0.3)
The MC truth of dx is 3.0 but I often get very different best-fit values after minimization. This is probably because my PDF, which has a non-zero bin width, is shifted and sum of three binned PDFs is used.
The best fit values obtained for each parameter are listed below:
MIGRAD MINIMIZATION HAS CONVERGED.
FCN=-73640.4 FROM MIGRAD STATUS=CONVERGED 264 CALLS 265 TOTAL
EDM=3.81233e-08 STRATEGY= 1 ERROR MATRIX UNCERTAINTY 0.0 per cent
EXT PARAMETER STEP FIRST
NO. NAME VALUE ERROR SIZE DERIVATIVE
1 dx 3.27545e+00 4.48015e-05 2.12661e-04 0.00000e+00
2 p0 7.21953e+03 8.77706e+01 -1.02551e-03 -2.00881e-02
3 p1 3.04091e+03 6.15363e+01 -6.97875e-04 -1.22533e-02
4 p2 1.83955e+03 4.52509e+01 4.18298e-04 -1.22703e-02
ERR DEF= 0.5
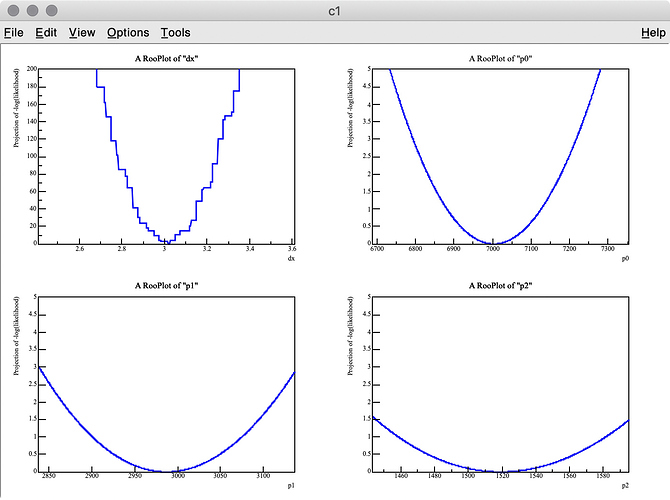
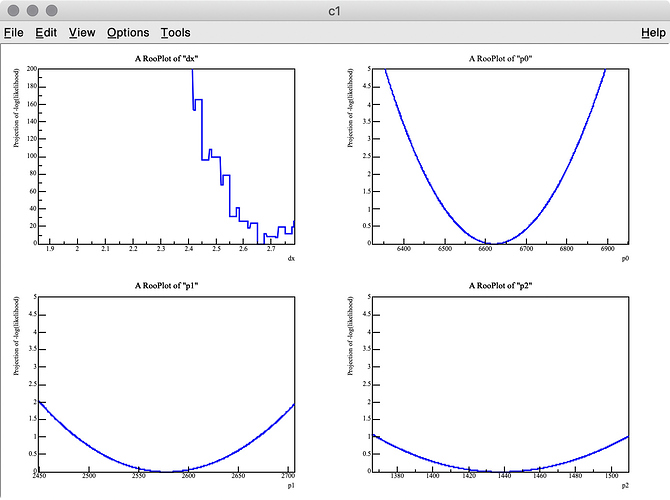
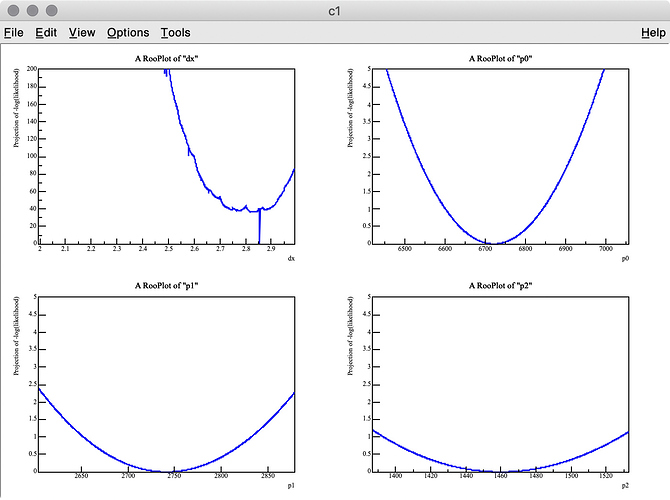
As you can see in the -log( likelihood ) of dx, it has spiky structure, resulting in wrong estimate of the best-fit value, whereas -log(likelihood) of p0, p1, and p2 are very smooth.
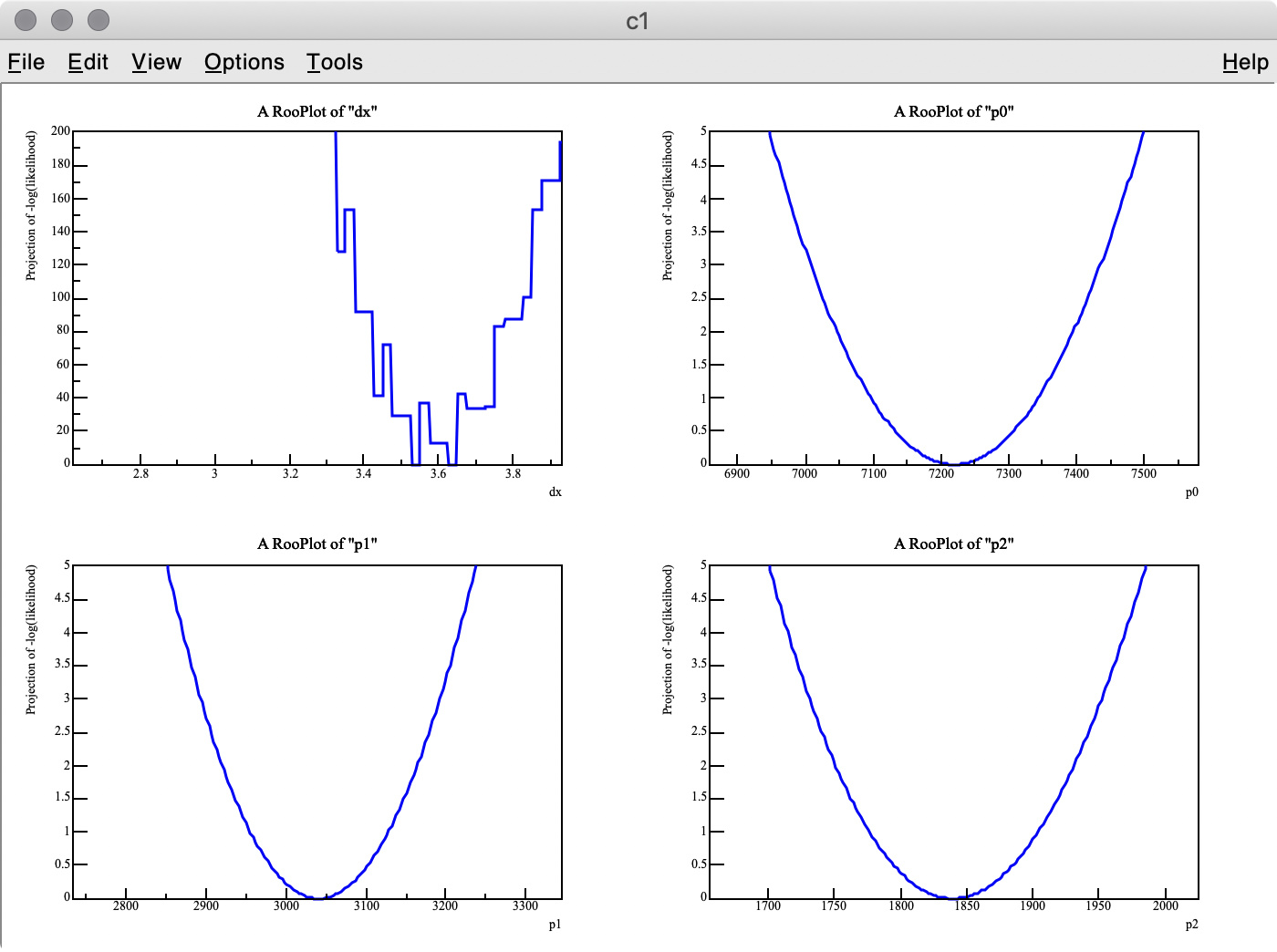
There are two points I am not certain about:
Q 1. Is there an easy way to find the best-fit dx value with the artificial peaks being ignored? I think a similar problem can happen when the energy scale of a MC PDF model is changed. I also tried to set the introder option to 1 to use interpolation, but it did not help much.
Q 2. I got 3.275 (MC truth = 3) as the best fit value of dx , but it looks like the minimum value of -log( likelihood ) of dx is around 3.6 in the top-left plot.
Why are these two values inconsitent?