I’m trying to do a fit across several MC samples, where most parameters remain constant across samples, but a few have a ~linear dependence on mass. I’m using a RooSimultaneous to do this, using the different MC mass points are my categories.
If I run on just one MC sample, everything seems to work fine - I get fits which fit the MC distributions well, and are correctly normalised.
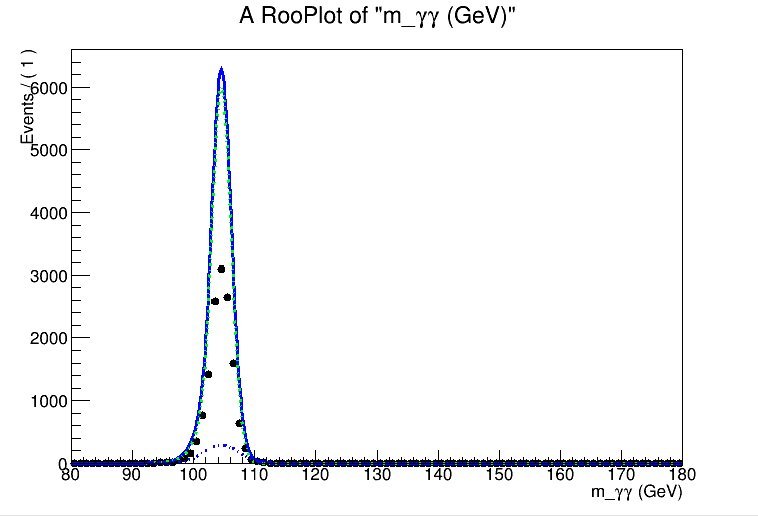
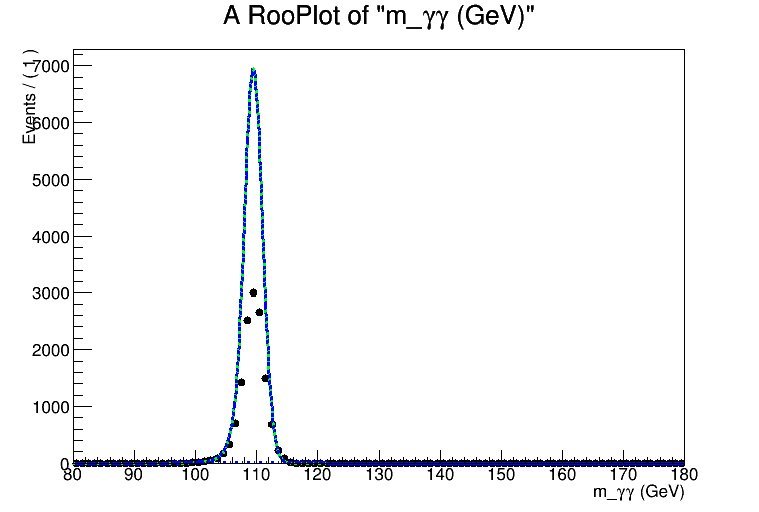
However, when I add further MC samples, the normalisation seems to go wonky - when I plot, it looks almost as though my individual pdfs are being normalised to the sum of the dataset, rather than only to the categories they are being fitted against. See the attachments - I’ve uploaded images of the result of performing this simultaneous fit to two.
I suspect this is a silly mistake on my part, since I’m not so experienced working with fits, but I’ve not had any luck trying to track down the reason. Does anybody know why my PDFs aren’t normalised properly?
I’ve included here the code of the method which creates and fits my pdfs (with most of the cruft removed - I left in the code where I produce the RooPlots, in case my issue is actually there):
void fit(int bin){
float sigyield, sigerr;
vector<RooDataSet*> RDS_ptbins;
RooDataSet* RDS_ptbin;
char sigcuts[200];
char invcuts[200];
char name[200];
char mname[100];
RooWorkspace *rws = new RooWorkspace("rws","rws");
std::map<std::string,RooDataSet*> _rdsmap;
RooCategory* index = new RooCategory("index","index");
RooSimultaneous *simpdf = new RooSimultaneous("simpdf","simpdf",*index);
for(int ii=0;ii<2; ii++){
RooWorkspace *wscat = new RooWorkspace("wscat","wscat"); //workspace for this category
int mass=105+(ii*5);
std::cout << "workspace for mass " << mass << std::endl;
sprintf(mname, "m%i", mass);
sprintf(sigcuts,"m_gg_truth == %i && pT > %f && pT < %f", mass, m_ptbounds.at(bin), m_ptbounds.at(bin+1));
RooDataSet* massbin = (RooDataSet*)RDS_mc->reduce(sigcuts);
index->defineType(mname);
_rdsmap[mname] = massbin;
//make vars
wscat->factory("mH["+TString(Form("%d",mass))+"]");
wscat->factory("m_gg[80,180]");
wscat->factory("mCBpar0[-0.550 , -25.0 , 25.0 ]"); //mean offset
wscat->factory("mCBpar1[0.0]"); //mean mass dependence
wscat->factory("aCBpar0[+1.550 , 0.0 , 10.0 ]"); //CB alpha
wscat->factory("aCBpar1[0.0]"); //alpha is kept constant across mass
wscat->factory("sCBpar0[+1.650 , 0.0 , 10.0]"); //CB sigma offset
wscat->factory("sCBpar1[+0.009 , -1.0 , 1.0]"); //CB sigma mass dependence
wscat->factory("kGApar0[+6.950 , 0.0 , 20.0 ]"); //gauss kappa
wscat->factory("fracCB[0.990,0.85,1.0]"); //CB fraction
wscat->factory("CB_n[10.0]"); //CB 'n' param kept constant at 10
//make pdf
wscat->factory("expr::CB_sigma('@1+(@0-125.0)*@2',mH,sCBpar0,sCBpar1)");
wscat->factory("prod::Gauss_sigma(kGApar0,CB_sigma)");
wscat->factory("expr::Gauss_mean('@0+@1+(@0-125.0)*@2',mH,mCBpar0,mCBpar1)");
wscat->factory("expr::CB_alpha('@1+(@0-125.0)*@2',mH,aCBpar0,aCBpar1)");
wscat->factory("RooGaussian::Gauss_Pdf(m_gg,Gauss_mean,Gauss_sigma)");
wscat->factory("RooCBShape::CB_Pdf(m_gg,Gauss_mean,CB_sigma,CB_alpha,CB_n)");
wscat->factory("SUM::CBGauss( fracCB*CB_Pdf,Gauss_Pdf )");
TString correlatedvars="m_gg,mCBpar0,mCBpar1,aCBpar0,aCBpar1,sCBpar0,sCBpar1,kGApar0,fracCB,CB_n";
//new workspace, with renamed vars to avoid conflicts
RooWorkspace *w = new RooWorkspace("w","w");
w->import((*wscat->pdf("CBGauss")),RenameAllNodes(mname),RenameAllVariablesExcept(mname,correlatedvars));
//add to main simpdf
simpdf->addPdf(*w->pdf(Form("CBGauss_%s",mname)),mname);
}
RooDataSet *combMC = new RooDataSet("combMC","combMC",*RAS_mc,Index(*index),Import(_rdsmap));
rws->import(*combMC);
rws->import(*simpdf);
rws->Print();
simpdf->fitTo(*combMC);
//now draw the fit results
for(int ii=0; ii<2; ii++){
int mass = 105+(5*ii);
sprintf(mname, "m%i", mass);
RooAbsPdf *thispdf = simpdf->getPdf(mname);
RooPlot* frame = V_m->frame();
combMC->plotOn(frame,Cut(Form("index==index::m%i",mass)));
thispdf->plotOn(frame);
thispdf->plotOn(frame,Components(Form("CB_Pdf_m%i",mass)),LineStyle(4),LineColor(3)) ;
thispdf->plotOn(frame,Components(Form("Gauss_Pdf_m%i",mass)),LineStyle(4),LineColor(4)) ;
sprintf(name,"m_H %i: %f < pT < %f",mass,m_ptbounds.at(bin),m_ptbounds.at(bin+1));
frame->SetNameTitle(name,name);
frame->Write();
}
return;
}
thanks
chris