Dear RooFit and Root experts,
When I do an extended maximum likelihood fit, I sometimes get that one of my parameters might have an asymmetric error which is 0 on the low side and X as big as the parameter for the error on the high side.
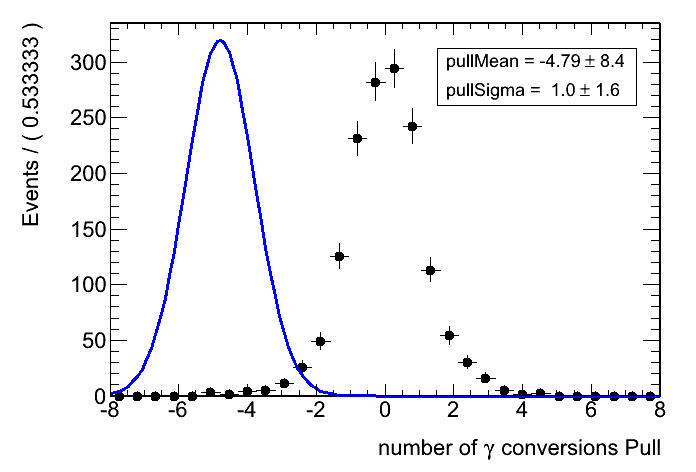
When I do pseudo-experiments on this case and I specify Minos errors, the pull distribution gets all mixed up:
[#0] WARNING:Minization -- RooFitGlue: Minimized function has error status.
Returning maximum FCN so far (-1e+30) to force MIGRAD to back out of this region. Error log follows
Parameter values: pullMean=4.79426, pullSigma=1
RooGenericPdf::pullGauss[ actualVars=(nconvpull,pullMean,pullSigma) formula="exp(-0.5*(@0-@1)*(@0-@1)/(@2*@2))" ]
getLogVal() top-level p.d.f evaluates to zero @ actualVars=(nconvpull = -inf,pullMean = 4.79426,pullSigma = 1)
getLogVal() top-level p.d.f evaluates to zero @ actualVars=(nconvpull = -inf,pullMean = 4.79426,pullSigma = 1)
getLogVal() top-level p.d.f evaluates to zero @ actualVars=(nconvpull = -inf,pullMean = 4.79426,pullSigma = 1)
getLogVal() top-level p.d.f evaluates to zero @ actualVars=(nconvpull = -inf,pullMean = 4.79426,pullSigma = 1)
getLogVal() top-level p.d.f evaluates to zero @ actualVars=(nconvpull = -inf,pullMean = 4.79426,pullSigma = 1)
getLogVal() top-level p.d.f evaluates to zero @ actualVars=(nconvpull = -inf,pullMean = 4.79426,pullSigma = 1)
getLogVal() top-level p.d.f evaluates to zero @ actualVars=(nconvpull = -inf,pullMean = 4.79426,pullSigma = 1)And the pull fit itself is shifted to the left (see image attached).
Is there a way to avoid this behaviour?
Many thanks in advance,
Clemencia
FMBtemplates_wPythiaB_4trackEtaBins_wTaus_new.root (24.2 KB)
rf801_mcstudy_MyModel_eta4_PythiaB.C (13.6 KB)