Dear All,
I am performing a simultaneous fit for signal strength mu. When I perform a fit with RooAbsPdf::createNll(), I often get a result that blows up with mu getting too largely negative. For example, as shown in these pre-fit and post-fit plots, and a standalone example at the bottom. Could anyone help explain what might cause this? Thanks very much.
Before the fit:
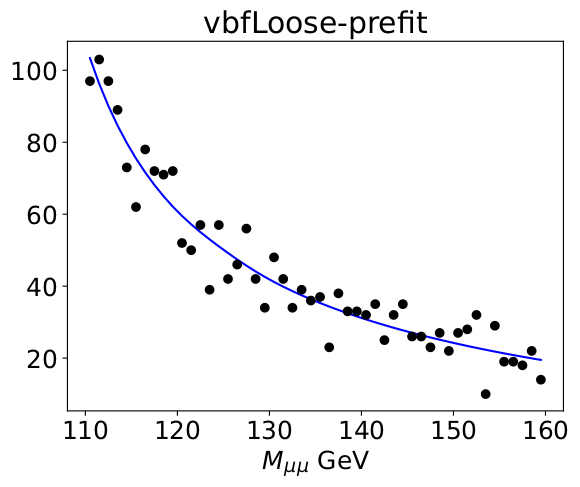
After the fit
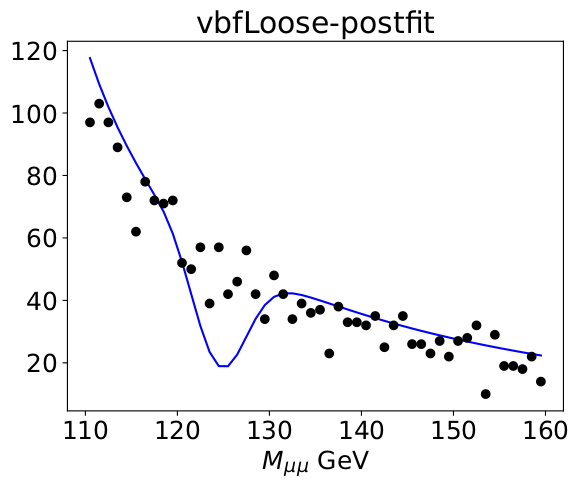
I have a standalone workspace with the pdf and roodatahist that I am using to make these attached.
Root file with workspace:
workspace.root (35.9 KB)
And a standalone script similar to how I am creating the NLL:
from __future__ import division
import sys, copy
sys.argv.append( '-b' )
from ROOT import TFile
from ROOT.RooFit import *
import ROOT
from ROOT import *
################################################################################
# standalone script for nll fit
################################################################################
# workspace
f = TFile.Open("workspace.root")
w = f.Get("w") # workspace
x = f.Get("x") # main variable
mu= f.Get("mu") # floating signal strength
# get rdh and pdf from workspace
# this is a RooDataHist with ONE "category" entry for simplicity
rdh = w.obj("simRdh")
# this is a RooSimultaneous with ONE category, and has ONE extended pdf added
pdf = w.obj("simPdf")
# create nll
nll = pdf.createNLL(rdh,RooFit.NumCPU(4))
ROOT.RooMinuit(nll).migrad()