Dear RooFit experts,
I am trying to use RooFit to fit some histogram with p.d.f created using RooHistPdf, but am getting some problems understanding the results. I observe what seems to me a discrepancy between what is plotted and the fit result. I’m doing a closure test, by fitting to the same histogram from which the p.d.f is derived. For simplicity, I consider only 2 bins.
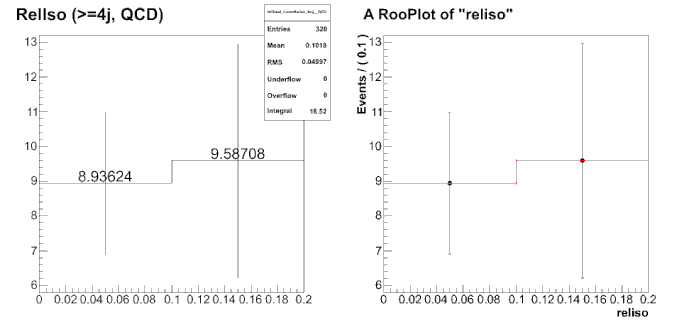
When I plot the p.d.f over the data points, I see perfect agreement. But when the fit results are slightly off by the true value. In my test, the true value is 18.5233 while the fit result is 18.0113.
So I wonder which is correct? The fit result or the plot?
I would be very grateful if you can help me to understand this discrepancy.
The plot I am referring to is shown below in the attachement (closure.gif).
The macro and the root file I am using are also attached.
Thank you very much.
Best wishes
Teh Lee
ps: I am using ROOT 5.22.00 and RooFit v2.95.
//
// 23-11-09
// Fit to all (s+b): Plan B
//
// - Use control sample as template.
// - Then fit to signal region
//
// closure test: take signal region template and fit it back
//-----------------------------
#include <iomanip>
using namespace RooFit;
void fit_closure(){
gStyle->SetOptStat(1111111);
double fit_from = 0;
double fit_upto = 0.2;
TFile f("test.root");
TCanvas c0("c0","fit control region",2*550,1*550);
c0.Divide(2,1);
c0.cd(1);
const double tiny = 1e-6;
///------------------------------------
/// Get template from control sample
///------------------------------------
TH1F *h = (TH1F*)f.Get("QCD_estimation/QCDest_CombRelIso_4mj__QCD");
h->Rebin(10);
h->GetXaxis()->SetRangeUser(0,fit_upto);
TH1F *control = h->Clone();
double ntrue = control->Integral(1,2);
cout << "sum of bin 1 and 2 = " << ntrue << endl;
control->GetXaxis()->SetRangeUser( 0, 0.2-tiny );
control->SetMarkerSize(2.5);
control->GetYaxis()->SetRangeUser(5.8,13.2);
control->DrawCopy("h text0");
// create HistPdf
RooRealVar reliso("reliso","reliso", 0, fit_upto);
reliso.setBins(10);
control->GetXaxis()->SetRangeUser(0,fit_upto);
RooDataHist rh_qcd("rh_qcd","qcd", reliso, control);
cout << "integral ori histo qcd = " << control->Integral() << endl;
RooHistPdf pdf_qcd( "pdf_qcd", "QCD pdf ", reliso, rh_qcd, 0);
// create model
RooRealVar nqcd("nqcd", "number of QCD events", 0, 100.0, "event");
RooAddPdf model("model","model", pdf_qcd, nqcd);
///--------------------------
/// Fit signal sample: "all"
///---------------------------
// Make a copy of the original histogram to fit
TH1F *all = (TH1F*)h->Clone();
// Prepare data in *fit range*
all->GetXaxis()->SetRangeUser(fit_from,fit_upto);
RooDataHist data("data","data in signal sample",reliso, all);
model.fitTo( data, Range( fit_from, fit_upto ) );
nqcd.Print();
RooPlot *isoframe = reliso.frame();
data.plotOn(isoframe, DataError(RooAbsData::SumW2), LineWidth(1));
model.plotOn(isoframe, LineColor(kRed), LineWidth(1));
c0.cd(2);
isoframe->SetAxisRange(5.8,13.2,"y");
isoframe->Draw();
c0.SaveAs( "closure.gif" );
}The results of the fit are shown below:
fit_closure.C (2.08 KB)
test.root (4.82 KB)