Hi @sanjeeda!
There are 3 problems I see in your scripts. Two I can help you fix immediately, and one I can only give general guidance for because I don’t know exactly how you expect your model to be.
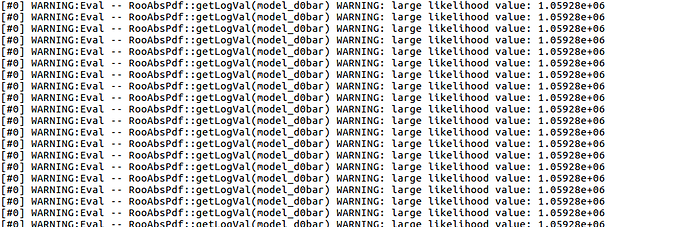
1st problem: you are ignoring the RooFit and Minuit warnings. If I run your script, I see some warnings about the sigma parameters for the Gaussians where you forgot to limit them in a positive range, and more warnings about limits that are too far apart for the yield variables. Fixing these usually makes the fit more stable. You should define a meaningful range for the sigma_ parameters, e.g.:
RooRealVar sigma_G1_sig("#sigma_{G1sig}", "sigma_gauss1",0.00385, 1e-5, 0.1);
// and not only:
// RooRealVar sigma_G1_sig("#sigma_{G1sig}", "sigma_gauss1",0.00385);
And for the yield parameters, I would narrow down the allowed order of magnitudes, e.g.:
RooRealVar sig_peak_yield("N_{sig_peak}","signal yield",3.18353e2,1e1,1e7);
// instead of
// RooRealVar sig_peak_yield("N_{sig_peak}","signal yield",3.18353e2,0,1e9);
// (note that allowing a zero yield is not a good idea anyway)
2nd problem: The RooFit variables in your combined dataset don’t match with the variables in the D0 and D0bar datasets.
At the beginning of the script, you defined six RooRealVar: mD0pi, mKK, mD0pi_d0, mKK_d0, mD0pi_d0bar, and mKK_d0bar. The variables with the _d0 and _d0bar suffixes are used to import the distributions_d0 and distributions_d0bar TTrees, which makes sense because if the variable names in the RooDataSet don’t match with the TTree branch names the branches don’t get imported.
But then you try to create a combined dataset with the variables without suffix:
RooDataSet combData("combData","combined data",RooArgSet(mKK, mD0pi),
Index(sample),
Import("D0",*dataxy_d0),Import("D0bar",*dataxy_d0bar)) ;
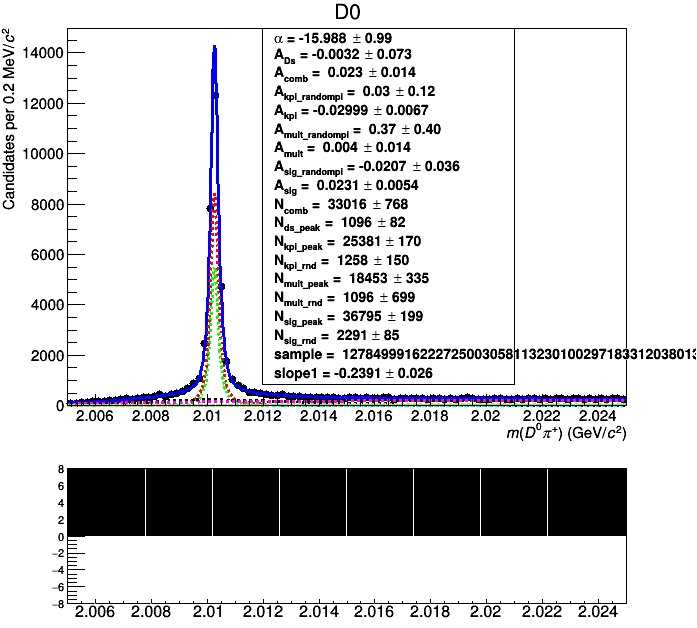
This does not work! The Import only works if the variable names match, otherwise the combined dataset will be just filled with the values that mKK and mD0pi are currently set to. If you comment out the line with simPdf.fitTo(combData) to skip the broken fit, you can indeed see that the data was not correctly imported.
You can work around this by replacing the RooDataSet constructor call before with some custom code that manually fills the dataset from the two existing ones even if the variable names don’t match:
RooArgSet dataVars{sample, mKK, mD0pi};
RooDataSet combData("combData","combined data",dataVars);
auto copyVal = [](RooAbsArg& to, RooAbsArg const& from){
static_cast<RooRealVar&>(to).setVal(
static_cast<RooAbsReal const&>(from).getVal());
};
sample.setLabel("D0");
for(std::size_t i = 0; i < dataxy_d0->numEntries(); ++i) {
RooArgSet const& vars = *dataxy_d0->get(i);
copyVal(dataVars["mkk"], vars["mkk_d0"]);
copyVal(dataVars["md0pi"], vars["md0pi_d0"]);
combData.add(dataVars);
}
sample.setLabel("D0bar");
for(std::size_t i = 0; i < dataxy_d0bar->numEntries(); ++i) {
RooArgSet const& vars = *dataxy_d0bar->get(i);
copyVal(dataVars["mkk"], vars["mkk_d0bar"]);
copyVal(dataVars["md0pi"], vars["md0pi_d0bar"]);
combData.add(dataVars);
}
I would also advise to put this code in a separate function to avoid spaghetti code. If you keep the fitTo call commented out and run your script again, you will see that the dataset is now filled correctly.
3rd problem (related to second one): you are using the wrong observable RooRealVars in your model.
As we discussed before, the only observables in the dataset are now mD0pi, mKK, plus a third variable sample that determines whether a given row is from a D0 or D0bar. This is the correct way to set up your dataset for the RooSimultaneous, but that also means that you can’t use the variables suffixed with _d0 or _d0bar in the model at all! They were only intermediate variables to import your trees separately! But you are using them in your model, and that’s where the evaluation errors come from.
So the 3rd change you need to make is to change your models to use only the mD0pi and mKK observables. However, I can’t tell you exactly how to do that, because from your script it’s not clear which parameters of the model should be shared between the D0 and D0bar components.
I see your final RooAddPdf models here:
RooAddPdf model_d0("model_d0","",
RooArgList(signal_peak,signal_rnd,
mult_peak,mult_rnd,kpi_peak, kpi_rnd,ds_peak, comb_bkg),
RooArgList(sig_peak_yield_d0, sig_randompi_yield_d0,
mult_peak_yield_d0, mult_randompi_yield_d0,
kpi_peak_yield_d0, kpi_randompi_yield_d0,d s_peak_yield_d0,
comb_yield_d0));
RooAddPdf model_d0bar("model_d0bar","",
RooArgList(signal_peak,signal_rnd,
mult_peak,mult_rnd,kpi_peak, kpi_rnd,ds_peak, comb_bkg),
RooArgList(sig_peak_yield_d0bar, sig_randompi_yield_d0bar,
mult_peak_yield_d0bar, mult_randompi_yield_d0bar,
kpi_peak_yield_d0bar, kpi_randompi_yield_d0bar,
ds_peak_yield_d0bar, comb_yield_d0bar));
I see that you have independent yield parameters for d0 and d0bar, which makes complete sense. But the RooAddPdf pdf components are exactly the same! Okay why not, but then I see comments in the pdf definitions like random pion (common for all), and sometimes you use the suffixed observables and sometimes not. That’s a bit fishy to me. Which pdfs/parameters should actually be shared then?
To conclude: I advise you to implement my suggestions to fix problem 1 and 2 first. Then you rewrite your model to not use the suffixed observables, carefully thinking which parameters and pdfs need to be shared between the D0 and D0bar components. Don’t hesitate to follow up here if you need help with this, but if you do please tell me what you really want to achieve mathematically as this is not clear to me.
Good luck!
Jonas
 .
.