Hi everyone,
I keep having this issue with normalization of extended likelihood fits (I am currently using pyRoot).
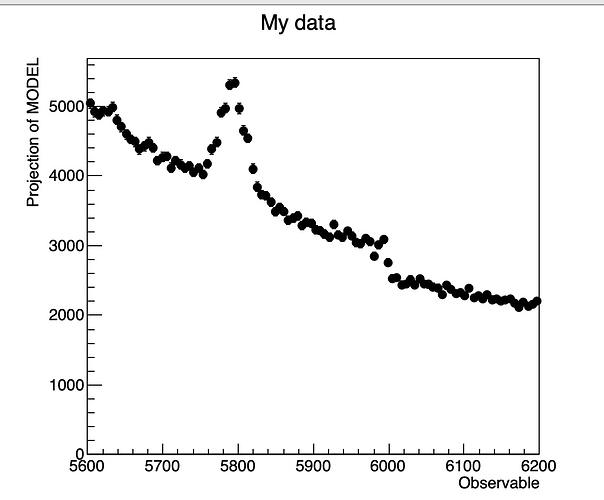
I am trying to fit binned data from a histogram with a composite model signal+background, but even if use RooExtendPdf when I plot model and dataset on the same canvas, the fit is normalized to 1 while data is obviously not.
I followed the user manual “Example 7” and double checked multiple times, but the problem won’t go away.
Here’s my code:
from ROOT import *
filename = TFile("../All_tmp.root")
tree = filename.Get("Xib0/mytree")
x = RooRealVar("lab0_M","Observable",5600,6200)
hist = TH1F("hist_name","hist title",100,5600,6200)
for event in tree:
hist.Fill(tree.lab0_M)
data = RooDataHist("data","my dataset",x,hist)
mean = RooRealVar("mean","mass mean",5800,5900,6200)
width = RooRealVar("width","gaussian width", 130,10,500)
gauss = RooGaussian("gauss","gaussian pdf",x,mean,width)
par1 = RooRealVar("param1","bgparam1",0.,100.,1.)
par2 = RooRealVar("param2","bgparam2",0.,100.,1.)
ground=RooArgusBG("bkg","background p.d.f.",x,par1,par2)
nsig = RooRealVar("nsig","signal fraction",10000,0.,80000.)
nbkg = RooRealVar("nbkg","background fraction",10000,0.,80000.)
Egaus = RooExtendPdf("esig","esig",gauss,nsig)
Ebkg = RooExtendPdf("ebkg","ebkg",ground,nbkg)
model = RooAddPdf("model","MODEL", RooArgList(Egaus,Ebkg))
xframe = x.frame(RooFit.Title("My data"))
model.plotOn(xframe)
data.plotOn(xframe)
model.fitTo(data)
xframe.Draw()
Can anyone help me?
Thanks a lot,
Andrea