Dear all,
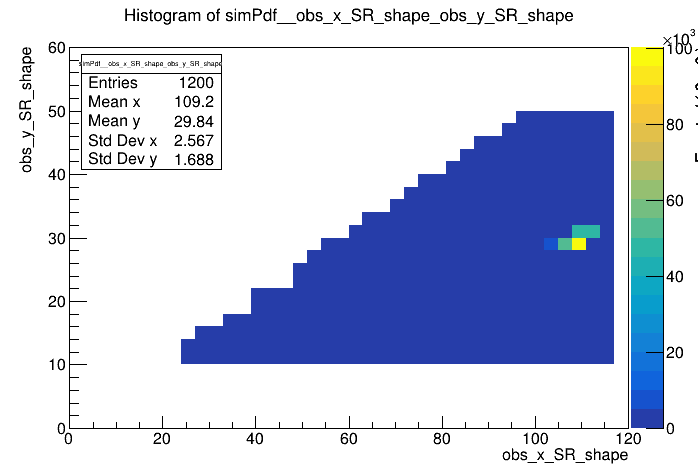
I created a workspace in Roofit from 2D histogram as in this example here:
#include "RooStats/HistFactory/MakeModelAndMeasurementsFast.h"
#include "TH2D.h"
void ShapeFit_diallo()
{
//Histos were already made by ShapeFit_makeHists.C. Let's open them up and use them
TFile* file_signal = new TFile("hists/BRscaled_gaussiansignal_allchannel_mZd30_mH110GeV.root");
TFile* file_data_bkg = new TFile("hists/Data_bkg_2D_all_channel_3-2GeVBin_interpNegBin.root");
//TFile* file_data = new TFile("hists/data_allchannel.root");
//signal
TH2D* h_sig = (TH2D*)file_signal->Get("Nominal/sig");
//TH2D* h_sig_scale_down = (TH2D*)file_bkg->Get("Nominal/bkg");
//TH2D* h_sig_scale_up = (TH2D*)file->Get("Signal_scale_up");
//background
TH2D* h_bkg = (TH2D*)file_data_bkg->Get("bkg_all");
//TH2D* h_bkg_slope_down = (TH2D*)file->Get("Background_slope_down");
//TH2D* h_bkg_slope_up = (TH2D*)file->Get("Background_slope_up");
//data
TH2D* h_obs = (TH2D*)file_data_bkg->Get("data_all");
//--------------------
//create the histfactory model
RooStats::HistFactory::Measurement meas("ShapeFit", "ShapeFit");
meas.SetOutputFilePrefix("workspaces/ShapeFit/2D");
meas.SetExportOnly(1);
meas.SetPOI("mu");
// this scales the histogram content, which already includes lumi, so set to 1
meas.SetLumi(1.0);
meas.SetLumiRelErr(0.02);
//create the parameters for the model
//first the signal normalization
RooStats::HistFactory::NormFactor normS;
normS.SetName("mu");
normS.SetHigh(100); // maximum value it can take
normS.SetLow(0); // minimum value it can take
normS.SetVal(1); // nominal value
//create the SR
RooStats::HistFactory::Channel SR("SR_shape");
SR.SetData(h_obs);
//add the signal and background samples
RooStats::HistFactory::Sample sample_S("S_shape");
sample_S.SetHisto(h_sig);
sample_S.AddNormFactor(normS);
//add the shape uncertainty for the signal
//RooStats::HistFactory::HistoSys shapeSys_S("Scale");
//shapeSys_S.SetHistoHigh(h_sig_scale_up);
//shapeSys_S.SetHistoLow(h_sig_scale_down);
//sample_S.AddHistoSys(shapeSys_S);
//let's also add a normalization systematic
//sample_S.AddOverallSys("sys_S",1./1.05,1.05);
SR.AddSample(sample_S);
RooStats::HistFactory::Sample sample_B("B_shape");
sample_B.SetHisto(h_bkg);
//add the shape uncertainty for the signal
//RooStats::HistFactory::HistoSys shapeSys_B("Slope");
//shapeSys_B.SetHistoHigh(h_bkg_slope_up);
//shapeSys_B.SetHistoLow(h_bkg_slope_down);
//sample_B.AddHistoSys(shapeSys_B);
//also add a normalization uncertainty
//sample_B.AddOverallSys("sys_B",1./1.15,1.15);
//SR.AddSample(sample_B);
//add the single region to the measurement
meas.AddChannel(SR);
//make the workspace
RooStats::HistFactory::MakeModelAndMeasurementFast(meas);
}
Then I wanted to compute the p0 value using StandardHypostestInv from this: ROOT: tutorials/roostats/StandardHypoTestInvDemo.C File Reference . I copy the macro and gives it my workspace as input but when I try to run it nothing happen. I attached my workspace.
Could someone tell me please what I am missing?
Thanks
Best,
Diallo.
2D_combined_ShapeFit_model.root (54.3 KB)