Hi.
First of all, I’m not good at coding.
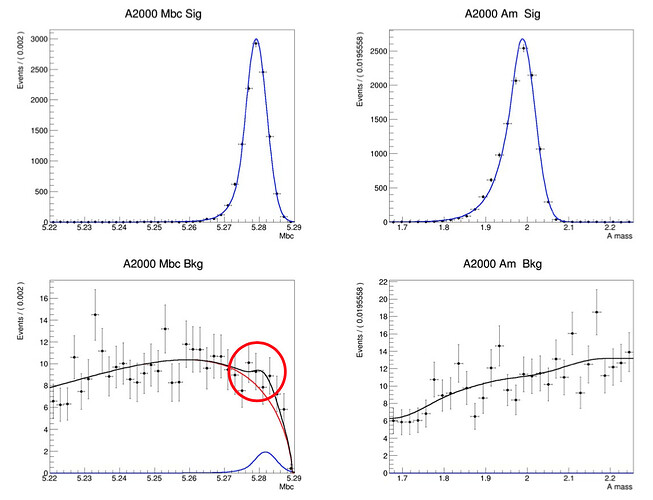
I want to do a 2D fit like below.
But I’ve only tried 1D fit, so I don’t know what to do with 2D fit.
As you can see in the bottom left of the picture, I want to do a 2D fit.
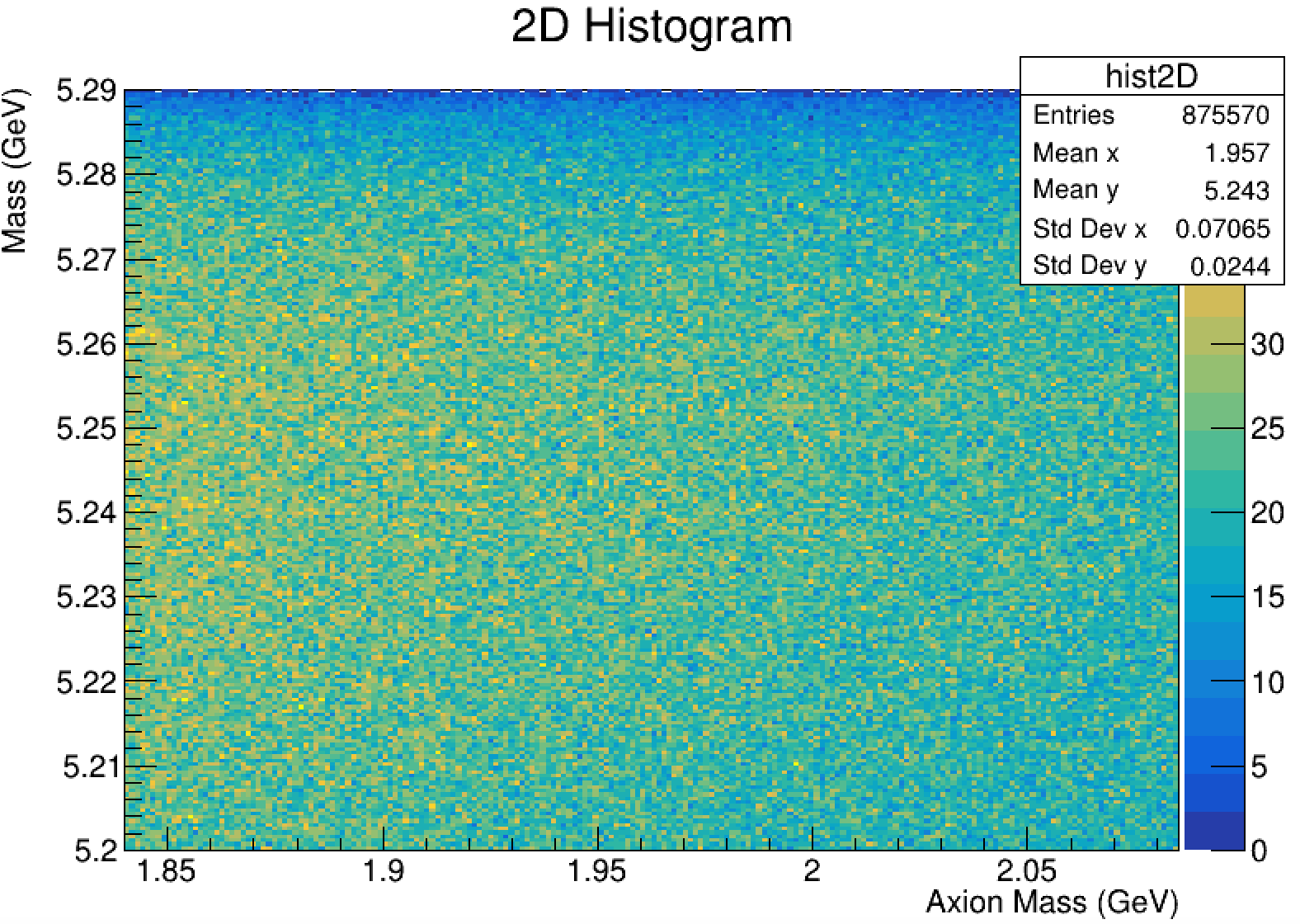
The pictures below are the 1D fitting and 2D histograms I did, something is off.
Finally, below is the code I was modifying.
I want to use KDE on the x-axis and argus+crystalball on the y-axis, in 2D fit.
That is, it should be (argus+crystalball)*KDE.
Can you tell me which part of my code I should fix?
Thank you for reading.
#include "RooRealVar.h"
#include "RooDataSet.h"
#include "RooGaussian.h"
#include "TCanvas.h"
#include "RooPlot.h"
#include "TAxis.h"
using namespace RooFit;
void gen_mbc()
{
TChain chain1("B_tree");
auto run_N = std::to_string(2);
auto file_2 = "/home/belle2/hyuna/MVA_ntuple/MVA_cc.root";
auto file_3 = "/home/belle2/hyuna/MVA_ntuple/MVA_uu.root";
auto file_4 = "/home/belle2/hyuna/MVA_ntuple/MVA_dd.root";
auto file_5 = "/home/belle2/hyuna/MVA_ntuple/MVA_ss.root";
auto file_6 = "/home/belle2/hyuna/MVA_ntuple/MVA_charged.root";
auto file_7 = "/home/belle2/hyuna/MVA_ntuple/MVA_mixed.root";
chain1.Add(file_2);
chain1.Add(file_3);
chain1.Add(file_4);
chain1.Add(file_5);
chain1.Add(file_6);
chain1.Add(file_7);
ROOT::EnableImplicitMT();
ROOT::RDataFrame df1(chain1);
TCut mvaCut = "outMVA < 0.17";
auto data_2D = new TH2F("y Axion Mass x Mbc",200,5.2,5.29,200,1.84,2.085);
chain1.Draw("B_axion_M:Mbc >> data_2D",mvaCut);
TH2F * hh = (TH2F *) gDirectory->Get("data_2D");
// auto df_m = df1.Filter(mvaCut.GetTitle());
// auto h = df_m.Histo1D({"Mbc", "Mbc", 200u, 5.2, 5.29}, "Mbc");
// auto h2 = df_m.Histo1D({"B_axion_M", "B_axion_M", 200u, 1.84, 2.085}, "B_axion_M");
RooRealVar x("Mass", "Mass(GeV)", 5.2, 5.29);
RooRealVar y("axion_Mass", "Axion mass(GeV)", 1.84, 2.085);
auto rd2h = RooDataHist("rd2h","rd2h",RooArgList(x,y),hh);
// auto rdh = RooDataHist("rdh","rdh",RooArgList(x),h.GetPtr(),1.0);
// auto rdh_2 = RooDataHist("rdh_2","rdh_2",RooArgList(y),);
// RooHistPdf m("Mbc", "", x, rdh, 0);
// RooHistPdf am("axion_Mass", "", y, rdh2, 0);
// Crystal ball PDF
RooRealVar m1("m1", "mean1 of crystalball", 5.27942, 5.27942, 5.27942);
RooRealVar s1("s1", "width1 of crystalball", 0.00315, 0.00315, 0.00315);
RooRealVar a1("a1", "alpha1 of crystalball", 1.32, 1.32, 1.32);
RooRealVar n1("n1", "n1 of crystalball", 10, 10, 10);
// Argus PDF
RooRealVar c("c", "slope parameter of argus", -1., -100., 100.);
RooCrystalBall crystal("crystal", "crystalball - signal", x, RooConst(5.279), s1, a1, n1);
RooArgusBG argus("argus", "argus - bkg", x, RooConst(5.29), c);
RooRealVar Nsig("Nsig", "yield of signal", 0.0, 0.0,10000000);
RooRealVar Nbkg("Nbkg", "yield of background", 0.0, 0.0, 10000000);
RooAddPdf model("model", "", RooArgList(crystal, argus), RooArgList(Nsig,Nbkg));
model.fitTo(rdh,Save());
// Fit pdf to data
RooFitResult* fitres = model.fitTo(rdh, RooFit::SumW2Error(true), Save());
RooArgSet fitargs = fitres->floatParsFinal();
TIterator* iter(fitargs.createIterator());
for (TObject* Tobj = iter->Next(); Tobj != 0; Tobj = iter->Next()) {
RooRealVar* rrv = dynamic_cast<RooRealVar*>(Tobj);
std::string name = rrv->GetName();
double val = rrv->getVal();
double err = rrv->getError();
printf("%s: %lf +- %lf\n", name.c_str(), val, err);
}
// Print values of mean and sigma (that now reflect fitted values and errors)
auto C = new TCanvas();
C -> cd();
h -> Draw();
C -> Update();
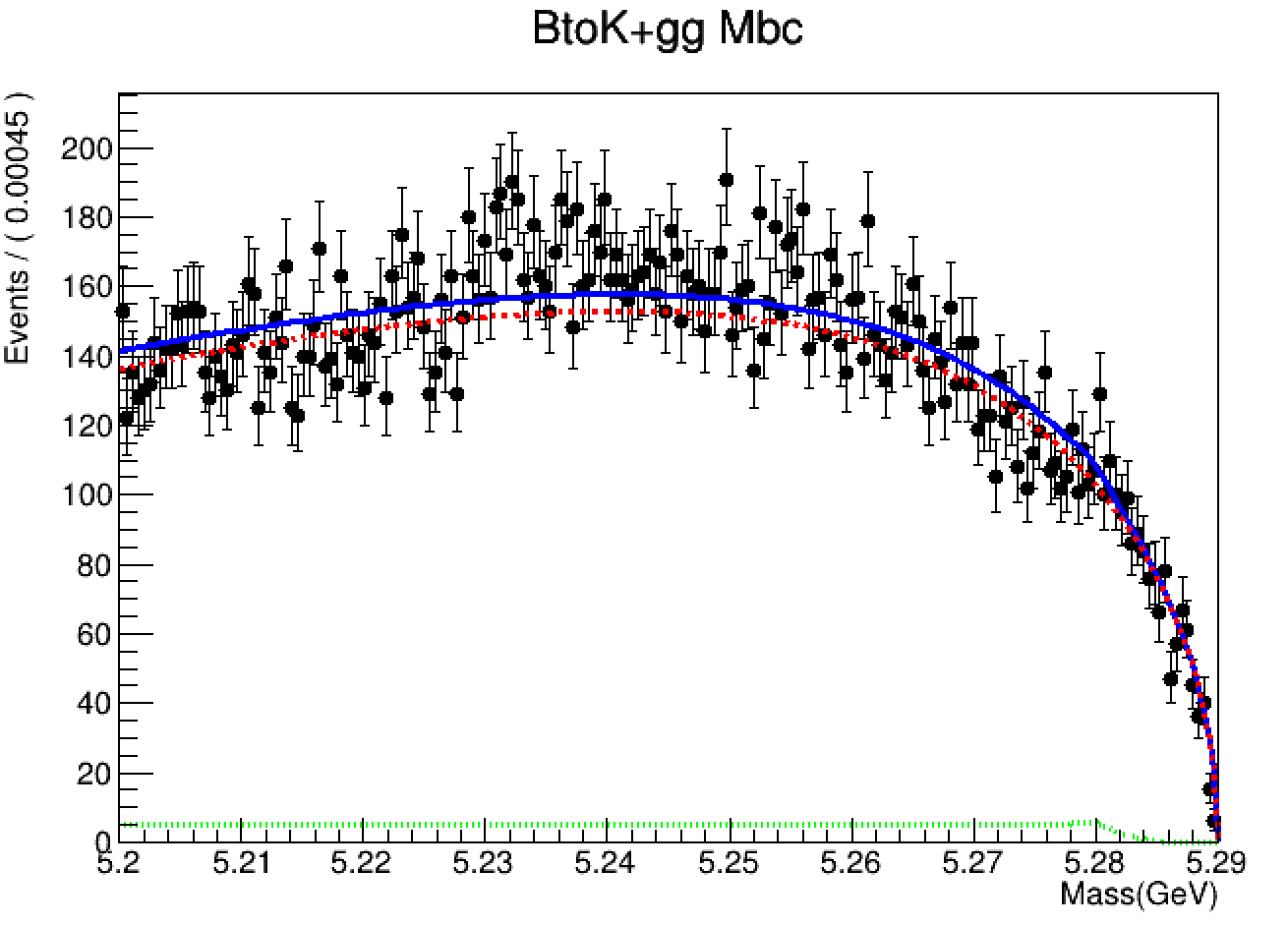
RooPlot* xframe = x.frame(Title("BtoK+gg Mbc"));
rdh.plotOn(xframe);
model.plotOn(xframe);
model.plotOn(xframe, Components(crystal), LineColor(kGreen), LineStyle(kDotted));
model.plotOn(xframe, Components(argus), LineColor(kRed), LineStyle(kDashed));
// model.paramOn(xframe, Label(""), Format("NEU", AutoPrecision(1)), Layout(0.1, 0.45, 0.9), Parameters(RooArgSet(Nsig,Nbkg,m1,s1,a1,n1,c)));
xframe->Draw();
C->SaveAs("2D_Mbc.png");