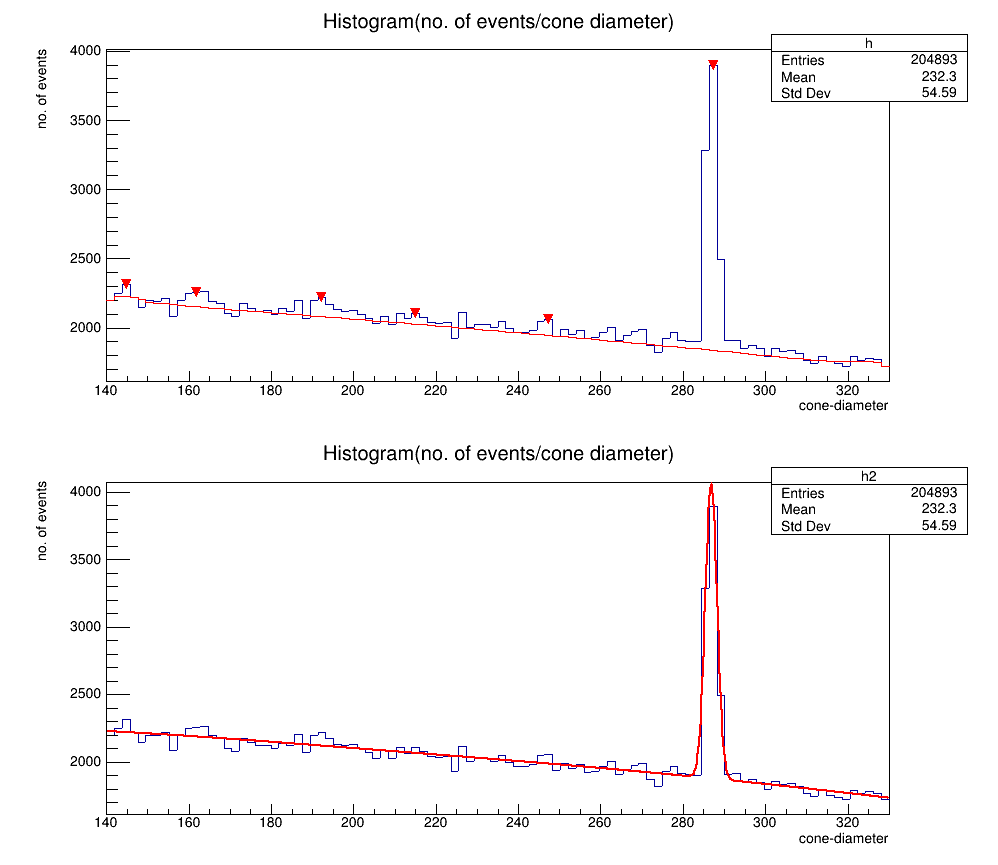
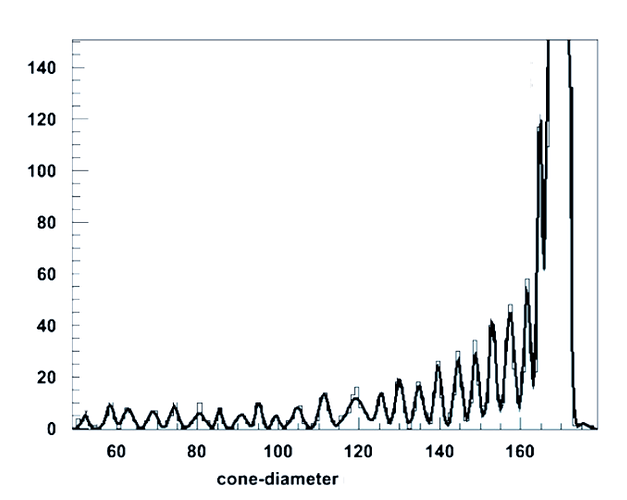
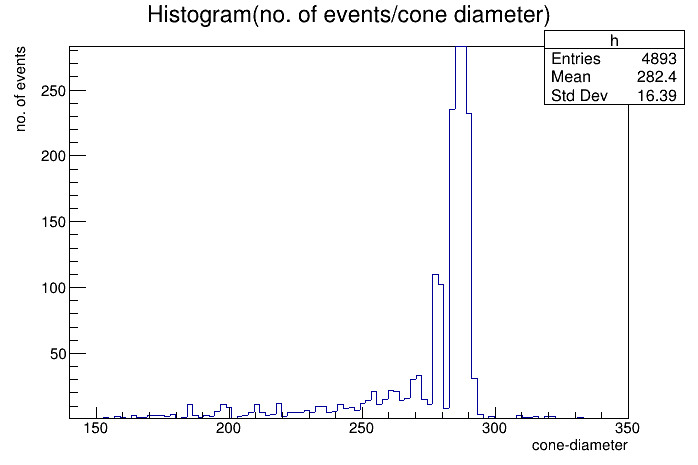
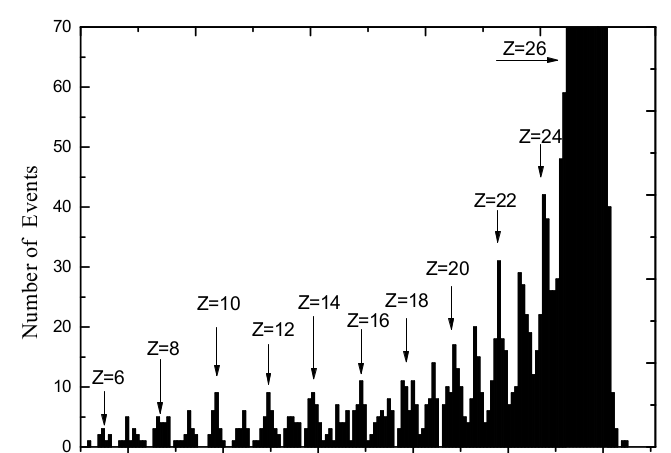
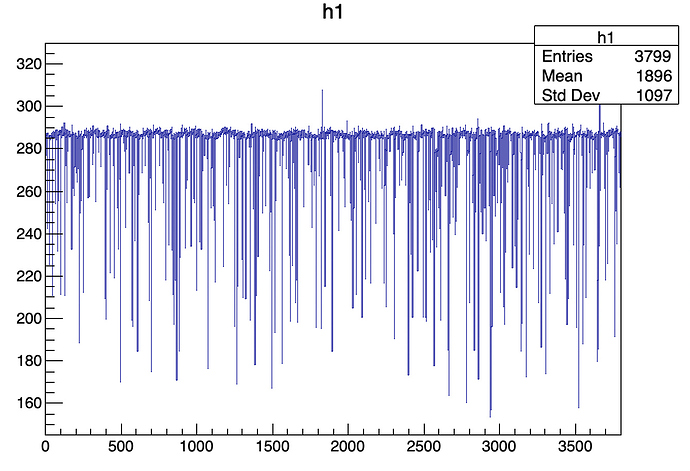
No, this example is for smoothing(which can be performed using draw panel) but i want to fit the Gaussian peak.
Here is the old example (which is not working in my system with root6 in ubuntu) for older root vs. in window system, can you tell me how this can be modified in my case :
// NPEAKS gaussian peaks + background (Gaus,Linear or Parabolic): 1-dimensional fit
// author: Gabriele Sirri
#include <iostream>
#include <fstream>
#define NPEAKS 9// <-- Number of peaks
#define MLIM 50 // Mean value limit of the signal fit ( m-MLIM < m < m+MLIM ) ADDITIONAL
#define SLIM 1.5 // Sigma limit of the signal fit ( s/SLIM < s < s*SLIM ) PROPORTIONAL
#define CLIM 5 // Constant limit of the signal fit ( c/CLIM < c < c*CLIM ) PROPORTIONAL
//#define NBCKPARS 3 // GAUSSIAN OR PARABOLIC BACKGROUND
//#define NBCKPARS 2 // LINEAR OR LANDAU BACKGROUND
#define NBCKPARS 0 // NO BACKGROUND
#define NPARS NPEAKS*3 + NBCKPARS
Double_t Pars[NPARS] ;
Double_t fitPars[NPARS] ;
//_____________________________________________________________________________
// The signal function: a gaussian
Double_t Signal(Double_t *x, Double_t *par)
{
Double_t arg = 0;
if (par[1]) arg = (x[0] - par[0])/par[1];
Double_t sig = par[2]*TMath::Exp(-0.5*arg*arg);
return sig;
// return TMath::Gaus(x[0],par[0],par[1],par[2]); // doesn't work...
}
// background function
Double_t Background(Double_t *x, Double_t *par)
{
//return TMath::Landau(x[0],par[0],par[1]); //Landau background // doesn't work...
//return par[0] + par[1]*x[0] ; //Linear background
//return par[0] + par[1]*x[0] + par[2]*x[0]*x[0]; //Quadratic background
//return Signal(x,par) ; //Gaussian background
return 0 ; // No Background
}
// Sum of NPEAKS peaks function + background
Double_t fitFunctionNB(Double_t *x, Double_t *par)
{
Double_t tot = 0 ;
Int_t a=0 ; // a= address of parameters array
for(a;a<NPEAKS;a++)
{
tot += Signal(x,&par[a*3]) ;
}
tot += Background(x,&par[a*3]) ;
return tot;
}
//_____________________________________________________________________________
// FIRST GUESS
void ReadPars(char* filename="a.txt")
{
ifstream f(filename);
Int_t i=0;
while(f>>Pars[i++] ) if(i==NPARS) break;
f.close();
}
void WritePars(char* filename="pars.txt")
{
ofstream f(filename);
for(Int_t i=0;i<NPARS;i++)
{
f<<fitPars[i];
if((i+1)%3) f<<" "; else f<<endl;
}
f.close();
}
//_____________________________________________________________________________
// Utilities
Double_t FindLowerLimit(TH1* h)
{
for(int k=0;k<h->GetNbinsX();k++)
if ( h->GetBinContent(k) )
return h->GetBinCenter(k);
}
Double_t FindUpperLimit(TH1* h)
{
for(int k=h->GetNbinsX()-1;k>-1;k--)
if ( h->GetBinContent(k) )
return h->GetBinCenter(k);
}
//_____________________________________________________________________________
TH1D* GetHistFromFile4(char* filename, Int_t nbinsx,Double_t xmin, Double_t xmax)
{
Double_t a,b;
Double_t pi=acos(-1);
Int_t c,d ;
TH1D* h1 = new TH1D("htemp","htemp",nbinsx,xmin,xmax);
ifstream f(filename);
while(f>>a>>b>>c>>d) {h1->Fill(a*b*pi); }
f.close();
h1->Draw();
return h1 ;
}
//_____________________________________________________________________________
TH1D* GetHistFromFile2(char* filename, Int_t nbinsx,Double_t xmin, Double_t xmax)
{
Double_t a,b,pi(acos(-1));
TH1D* h1 = new TH1D("htemp","htemp",nbinsx,xmin,xmax);
ifstream f(filename);
while(f>>a>>b) h1->Fill(a*b*pi);
f.close();
h1->Draw();
return h1 ;
}
//_____________________________________________________________________________
// *************************************************************************
// Fit NPEAKS gaussian peaks + background
void backsigNB(char* fitoption="V")
{
int npar = NPARS;
int bckpar = NBCKPARS;
int peakpar = 3;
Double_t x1 = FindLowerLimit(htemp);
Double_t x2 = FindUpperLimit(htemp);
printf("x1: %f\t x2: %f\n",x1,x2);
Double_t* guess_par = &Pars ;
// create a TF1 with the range from -1.1 to 1.1 and 24 parameters
TF1 *fitFcn = new TF1("fitFcn",fitFunctionNB,x1,x2,npar);
fitFcn->SetParameters(guess_par);
fitFcn->SetNpx(300);
// Set Limits for the signal parameters
for(int j=0;j<NPEAKS;j++)
{
int ipar = 3*j;
fitFcn->SetParName(ipar ,"Mean_value");
fitFcn->SetParName(ipar+1,"Sigma");
fitFcn->SetParName(ipar+2,"Constant");
//fitFcn->SetParLimits(ipar,50,1000);
Double_t mval = fitFcn->GetParameter(ipar);
fitFcn->SetParLimits(ipar ,mval-MLIM,mval+MLIM);// define parameters LIMITS for mean value
Double_t sval = fitFcn->GetParameter(ipar+1);
fitFcn->SetParLimits(ipar+1,sval/SLIM ,sval*SLIM);// define parameters LIMITS for sigma
// Double_t cval = fitFcn->GetParameter(ipar+2);
// fitFcn->SetParLimits(ipar+2,cval/CLIM ,cval*CLIM);// define parameters LIMITS for constant
}
htemp->Fit("fitFcn",fitoption);
Double_t BinWidth = htemp->GetBinWidth(0);
Double_t* fit_par = new Double_t[npar];
// writes the fit results into the par array
fitFcn->GetParameters(fit_par);
// PRINT RESULTS
// Peak Area Evaluation
Double_t signalarea;
for( j=0;j<NPEAKS;j++)
{
int ipar =3*j;
char funcname[16];
sprintf (funcname,"peakFcn%d",j);
TF1 *peakFcn = new TF1(funcname,Signal,x1,x2,peakpar);
peakFcn->SetParameters(&fit_par[ipar]);
Double_t peakarea = peakFcn->Integral(x1,x2) /BinWidth;
printf("Peak %2d Mean: %7.1f\tSigma: %5.1f\tConst: %7.1f Area:%11.3f\n", j,
fit_par[ipar] , fit_par[ipar+1] , fit_par[ipar+2] , peakarea );
signalarea += peakarea;
// improve the picture:
//peakFcn->SetLineColor(j%2?5:4); // comment this two lines to not plot single peaks
//peakFcn->Draw("same"); // comment this two lines to not plot single peaks
}
printf("\t\t\t\t\t\t\t\t----------\n");
printf("\t\t\t\t\t\t Total Peaks:%11.3f\n\n", signalarea);
// Background Evaluation
Double_t bckarea;
{
TF1 *bckFcn = new TF1("bckFcn",Background,x1,x2,bckpar);
bckFcn->SetParameters(&fit_par[NPEAKS*3]);
bckarea = bckFcn->Integral(x1,x2) /BinWidth;
printf("Background ");
for(Int_t i=0;i<NBCKPARS;i++)
printf("p%d:%f ",i,fit_par[NPEAKS*3+i] );
printf("\t Area:%11.3f\n", bckarea);
// improve the picture:
//bckFcn->SetLineColor(3); // comment this two lines to not plot background
//bckFcn->Draw("same"); // comment this two lines to not plot background
}
Double_t fitarea = fitFcn->Integral(x1,x2) /BinWidth;
printf("\t\t\t\t\t\t\t\t----------\n");
printf("\t\t\t\t\t\t Total Area:%11.3f\n\n", fitarea);
printf("\t\t\t\t\t\t Total Entries:%11.3f\n", htemp->Integral(x1,x2) );
printf("\t BIN WIDTH: %f\n",BinWidth) ;
fitFcn->Draw("same");
//EXPORT pars
for(Int_t i=0;i<NPARS;i++) fitPars[i]= fit_par[i];
}
//
// *************************************************************************
void Do()
{
h = GetHistFromFile4("a.txt",150,0,4500);
ReadPars();
backsigNB();
WritePars();
}