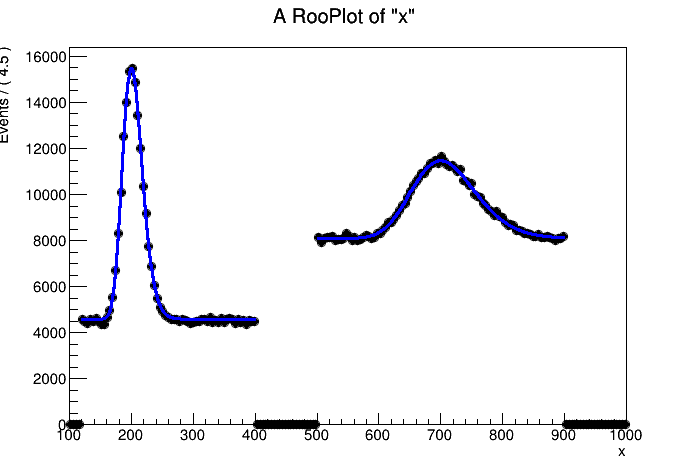
I have a spectrum, with 2 (or more) distant peaks. I want to fit all of them with a gaussian, where the sigma is constrained by a*energy + b, here a and b should be optimized simultaneously. And I want each of the peaks to have a separate constant background (i.e. a constant that is only nonzero in the vicinity of the peakregion).
For the moment, I use:
RooRealVar a_sig("a_sig","a of sigma relation", 0.00084, 0, 0.1,"keV");
RooRealVar b_sig("b_sig","b of sigma relation", 0.1, 0, 1,"keV");`
And then I make multiple ranges:
E.setRange(range_name.str().c_str(),downenergy, upenergy);
Then I make the background:
constback[ii] = new RooRealVar(constback_name.str().c_str()," ",
15, 0, 1000,"keV");
constback_touse[ii] =
new RooFormulaVar(constback_touse_name.str().c_str(),
"(@0 > (@2-20))*(@0 < (@3+20)) *@1",
{*mx_const[ii],*constback[ii], *Roodown[ii], *Rooup[ii]});
where Rooup and down is just boundaries for the where I want the background.
And I define my model for each of the peaks as:
Sig[ii] =
new RooGenericPdf(sig_name.str().c_str(),
"(exp(-((@0 - @1)/(@2)) * ((@0 - @1)/@2)/2) /(@2*2.5) + @3)",
{E,*mx[ii],*sigma[ii], *constback_touse[ii]});
eSig[ii] = new RooExtendPdf(eSig_name.str().c_str()," ",*Sig[ii],*nSig[ii]);
Then the fit is performed as:
RooArgList pdfList{"pdfList"};
for(int i = 0; i < numPeaks; i++) {
pdfList.add(*(eSig[i]));
}
RooAddPdf finalPDF("finalPDF","finalPDF",pdfList);
RooNLLVar nll("nll", "nll", finalPDF, data,
Range(all_ranges.str().c_str()), Extended(kTRUE), NumCPU(100));
RooMinimizer m(nll) ;
m.migrad() ;
m.hesse() ;
RooFitResult* r_sig = m.save() ;
I have 3 problems, the first one is, that when I use:
finalPDF.plotOn(frame, RooFit::NormRange(all_ranges.str().c_str()));
the background is clearly not used the way I want, I would expect a jump at Rooup[ii] +20, with Rooup = 855, but the background is just straight.
Moreover, I noticed that, the larger I make the interval for E (i.e. increasing the 1185 or decreasing the 830 in this definition: RooRealVar E("E","Energy1",830, 1185,"keV");), the worse the fit becomes. Even if I only fit one peak, it looks like less datapoints are taken into account?
Lastly, making this fit is taking >30min, while if I do not add the clipping, it is done in about a minute, which is rather weird to me.