Dear all,
I’ve been using Roofit for a long time now and have never spent too much time optimizing the performance of my fit function. With this post I would like to ask for recommendations and suggestions.
The problem:
Fitting a sum function (“hyperEMG”) of several exponentially-modified Gaussians (“EMG”) to a TH1D (the idea behind the hyperEMG is published here).
The EMG is defined as the convolution of a Gaussian with an exponential distribution, which has an analytical representation in the form of:
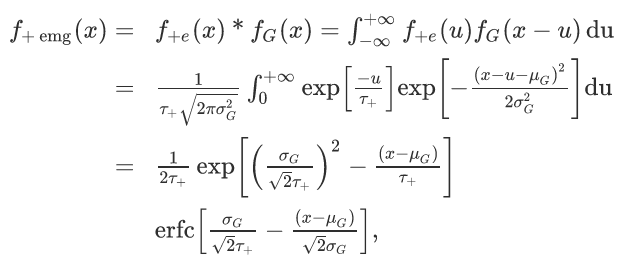
including the Gaussian mean and variance mu and sigma, and the decay constant tau from the exponential
The hyperEMG is the sum of n EMGs with negative decay constants and m EMGs with positive decay constants. For my data, the Gaussian mean and variance are shared between the different EMGs.
My approach
As I did not want to calculate the convolution, I implemented the analytical expression of the EMG. Each hyperEMG is built from a RooGenericPdf, which takes the analytical function as a string. Consider a hyperEMG with one negative and two positive components. In terms of RooRealVar we have one sigma, one mu, three tau, and two ratios to mix the three EMGs together. As a function string this looks like this
@4*1/(2*@3)*exp((@2/(1.4142*@3))^2+(@0-@1)/@3)*TMath::Erfc(@2/(1.4142*@3)+(@0-@1)/(1.4142*@2))+@6*1/(2*@5)*exp((@2/(1.4142*@5))^2-(@0-@1)/@5)*TMath::Erfc(@2/(1.4142*@5)-(@0-@1)/(1.4142*@2))+(1-@4-@6)*1/(2*@7)*exp((@2/(1.4142*@7))^2-(@0-@1)/@7)*TMath::Erfc(@2/(1.4142*@7)-(@0-@1)/(1.4142*@2))
where the RooRealVar are denoted with the @ symbol.
Now, this is for one species in my spectrum, which can consist of many species. The assumption is that all species share the same sigma and tau parameters, as well as the EMG mixing ratios, which means that those RooRealVar are used for all species. Now considering we use the abovementioned function for three different species, we will have to fit three mu, one sigma, three tau, two EMG mixing ratios, and two mixing ratios for the three species, resulting in 11 fit parameters. The mixing of the different species is eventually being done using RooAddPdf on the three individual RooGenericPdf.
This example is the most complex I could fit so far. Having to fit more species or having to use more EMG components results in a very inefficient fitting, even though the fit eventually converges.
The question
Without asking for an explicit implementation of this problem, how would you tackle this (strictly conceptually speaking)?
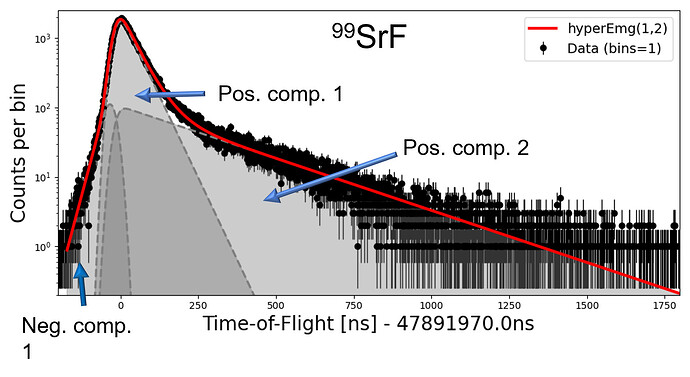
Finally, here is an example where this three-component hyperEMG is fitted to real data.
Thank you,
Lukas