Is there a method to perform a deconvolution in RooFit?
I used the RooFFTConvPdf function to convolve two RooHistPdf’s and was wondering if there was a reverse (deconvolution function) that would give me back the original input.
I am aware of the deconvolution in TSpectrum but that seems to deal specifically with peaks, which doesn’t work for my case.
Here are the specific lines of code I am discussing:
[code]
RooRealVar x(“x”,“x”, 0, 2500);
//create DataHist for pdf creation
RooDataHist* RDHh2 = new RooDataHist(“RDHh2”, " ", x, h2);
RooDataHist* RDHPulse = new RooDataHist(“RDHPulse”, " ", x, PulseHisto);
RooHistPdf* pdfh2 = new RooHistPdf(“pdfh2”, “pdfh2”, x, RDHh2, 2);
RooHistPdf pdfPulse = new RooHistPdf(“pdfPulse”, “pdfPulse”, x, *RDHPulse, 2);
//RooFFTConvPdf
RooFFTConvPdf expTOF(“ExpTOF”, “convolved spectrum”, x, *pdfh2, *pdfaccPulse);
TH1 *histoExpTOF = expTOF.createHistogram(“histoExpTOF”,x,0,0,0);
TH1 *histoPdfPulse = pdfPulse->createHistogram(“pdfPulse”,x,0,0,0);
}[/code]
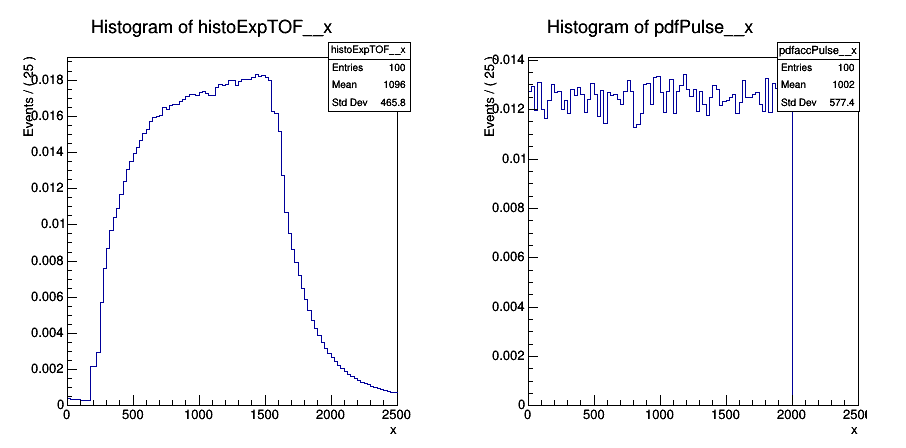
I am attempting to deconvolve the convolved histoExpTOF with histoPdfPulse, where histoExpTOF is convolved with histoPdfPusle. Here are figures of each histogram.