I’ve been trying to run chi2FitTo on histograms within a fitting range using the Range() option. This has worked for running a likelihood fit, but when using a chi2 fit, it has failed. I’m not sure if this is some interaction I haven’t accounted for, or just a quirk of using chi2FitTo, but some clarification would be welcome. I’ve tested with rf602_chi2fit.C and I would share but I’m a new user so can’t add links
Welcome to the ROOT Forum!
And I’m sure @jonas can help you
So rf602_chi2fit.C doesn’t work for you when you specify a range? You can post here the code (full macro or only the line/s that you changed), just copy & paste it but enclose the code in three single backquotes (“backticks”, i.e. three of this: ` without spaces), at the beginning and end of the block of lines of code, e.g. type this here in your post (without the backslashes!) and see what you get:
\```
my code line 1;
my code line 2;
// etc
\```
If I change this line in the example:
//model.chi2FitTo(*dh, PrintLevel(-1));
model.chi2FitTo(*dh, Range(0.,1.));
I get (with ROOT 6.36.10)
[#0] WARNING:InputArguments -- The parameter 'sigma1' with range [-inf, inf] of the RooGaussian 'sig1' exceeds the safe range of (0, inf). Advise to limit its range.
[#0] WARNING:InputArguments -- The parameter 'sigma2' with range [-inf, inf] of the RooGaussian 'sig2' exceeds the safe range of (0, inf). Advise to limit its range.
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: activating const optimization
[#1] INFO:Minimization -- The following expressions have been identified as constant and will be precalculated and cached: (sig1,sig2)
[#1] INFO:Minimization -- The following expressions will be evaluated in cache-and-track mode: (bkg)
Minuit2Minimizer: Minimize with max-calls 2000 convergence for edm < 1 strategy 1
Info in <Minuit2>: MnSeedGenerator Computing seed using NumericalGradient calculator
Info in <Minuit2>: MnSeedGenerator Evaluated function and gradient in 248.553 μs
Info in <Minuit2>: MnSeedGenerator Initial state: FCN = 106.1754789 Edm = 1.260070585 NCalls = 17
Info in <Minuit2>: MnSeedGenerator Initial state
Minimum value : 106.1754789
Edm : 1.260070585
Internal parameters: [ 0 -0.6435011088 0 0.6435011088]
Internal gradient : [ -9.229193845 35.69317399 24.70460566 9.713828671]
Internal covariance matrix:
[[ 0.0021104285 0 0 0]
[ 0 0.0034850852 0 0]
[ 0 0 0.0001669053 0]
[ 0 0 0 0.0033769849]]]
Info in <Minuit2>: VariableMetricBuilder Start iterating until Edm is < 0.002 with call limit = 2000
Info in <Minuit2>: VariableMetricBuilder 0 - FCN = 106.1754789 Edm = 1.260070585 NCalls = 17
Info in <Minuit2>: VariableMetricBuilder 1 - FCN = 105.1128131 Edm = 0.08526485556 NCalls = 27
Info in <Minuit2>: VariableMetricBuilder 2 - FCN = 104.86813 Edm = 0.1538210632 NCalls = 37
Info in <Minuit2>: VariableMetricBuilder 3 - FCN = 104.6396334 Edm = 0.0006219311146 NCalls = 47
Info in <Minuit2>: MnHesse Done after 240.277 μs
Info in <Minuit2>: VariableMetricBuilder After Hessian
Info in <Minuit2>: VariableMetricBuilder 4 - FCN = 104.6396334 Edm = 0.0007780570477 NCalls = 70
Info in <Minuit2>: VariableMetricBuilder Stop iterating after 605.568 μs
Minuit2Minimizer : Valid minimum - status = 0
FVAL = 104.639633447510988
Edm = 0.000778057047730882148
Nfcn = 70
a0 = 0.501526 +/- 0.0229096 (limited)
a1 = 0.158456 +/- 0.0368354 (limited)
bkgfrac = 0.506609 +/- 0.011349 (limited)
sig1frac = 0.815448 +/- 0.0373695 (limited)
Info in <Minuit2>: Minuit2Minimizer::Hesse Using max-calls 2000
Info in <Minuit2>: MnHesse Done after 232.272 μs
Info in <Minuit2>: Minuit2Minimizer::Hesse Hesse is valid - matrix is accurate
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: deactivating const optimization
90.865
So I’ve changed the macro to:
#include "RooRealVar.h"
#include "RooDataSet.h"
#include "RooGaussian.h"
#include "RooChebychev.h"
#include "RooAddPdf.h"
#include "TCanvas.h"
#include "TAxis.h"
#include "RooPlot.h"
using namespace RooFit;
void rf602_chi2fit()
{
// S e t u p m o d e l
// ---------------------
// Declare observable x
RooRealVar x("x", "x", 0, 10);
// Create two Gaussian PDFs g1(x,mean1,sigma) anf g2(x,mean2,sigma) and their parameters
RooRealVar mean("mean", "mean of gaussians", 5);
RooRealVar sigma1("sigma1", "width of gaussians", 0.5);
RooRealVar sigma2("sigma2", "width of gaussians", 1);
RooGaussian sig1("sig1", "Signal component 1", x, mean, sigma1);
RooGaussian sig2("sig2", "Signal component 2", x, mean, sigma2);
// Build Chebychev polynomial pdf
RooRealVar a0("a0", "a0", 0.5, 0., 1.);
RooRealVar a1("a1", "a1", 0.2, 0., 1.);
RooChebychev bkg("bkg", "Background", x, RooArgSet(a0, a1));
// Sum the signal components into a composite signal pdf
RooRealVar sig1frac("sig1frac", "fraction of component 1 in signal", 0.8, 0., 1.);
RooAddPdf sig("sig", "Signal", RooArgList(sig1, sig2), sig1frac);
// Sum the composite signal and background
RooRealVar bkgfrac("bkgfrac", "fraction of background", 0.5, 0., 1.);
RooAddPdf model("model", "g1+g2+a", RooArgList(bkg, sig), bkgfrac);
// C r e a t e b i n n e d d a t a s e t
// -----------------------------------------
std::unique_ptr<RooDataSet> d{model.generate(x, 10000)};
std::unique_ptr<RooDataHist> dh{d->binnedClone()};
// Construct a chi^2 of the data and the model.
// When a pdf is used in a chi^2 fit, the probability density scaled
// by the number of events in the dataset to obtain the fit function
// If model is an extended pdf, the expected number events is used
// instead of the observed number of events.
x.setRange("fitRange", 3, 7);
model.chi2FitTo(*dh, {PrintLevel(-1)}, Range("fitRange"));
// NB: It is also possible to fit a RooAbsReal function to a RooDataHist
// using chi2FitTo().
// Note that entries with zero bins are _not_ allowed
// for a proper chi^2 calculation and will give error
// messages
std::unique_ptr<RooAbsData> dsmall{d->reduce(EventRange(1, 100))};
std::unique_ptr<RooDataHist> dhsmall{static_cast<RooDataSet&>(*dsmall).binnedClone()};
std::unique_ptr<RooAbsReal> chi2_lowstat{model.createChi2(*dhsmall)};
cout << chi2_lowstat->getVal() << endl;
TCanvas *cdata = new TCanvas("sPlot", "sPlot demo", 400, 600);
RooPlot *frame = x.frame(Title("Fit of model to discriminating variable"));
dh->plotOn(frame);
model.plotOn(frame, Name("FullModel"));
frame->Draw();
}
The final frame drawing is just to show the fit isn’t limited, but the main change is adding a named range. Using number limits doesn’t changed the output though. And it gives:
Processing rf602_chi2fit.C…
[#1] INFO:Eval – RooRealVar::setRange(x) new range named ‘fitRange’ created with bounds [3,7]
[#0] WARNING:InputArguments – The parameter ‘sigma1’ with range [-inf, inf] of the RooGaussian ‘sig1’ exceeds the safe range of (0, inf). Advise to limit its range.
[#0] WARNING:InputArguments – The parameter ‘sigma2’ with range [-inf, inf] of the RooGaussian ‘sig2’ exceeds the safe range of (0, inf). Advise to limit its range.
[#1] INFO:Minimization – RooAbsMinimizerFcn::setOptimizeConst: activating const optimization
[#1] INFO:Minimization – The following expressions have been identified as constant and will be precalculated and cached: (sig1,sig2)
[#1] INFO:Minimization – The following expressions will be evaluated in cache-and-track mode: (bkg)
[#1] INFO:Minimization – RooAbsMinimizerFcn::setOptimizeConst: deactivating const optimization
90.865
With the below saved as fit2.C and run with
for i in 0 1; do root -l -b -q "fit2.C(0,3,$i)"; root -l -b -q "fit2.C(3,7,$i)"; root -l -b -q "fit2.C(7,10,$i)"; done
#include "RooRealVar.h"
#include "RooDataSet.h"
#include "RooGaussian.h"
#include "RooChebychev.h"
#include "RooAddPdf.h"
#include "TCanvas.h"
#include "TAxis.h"
#include "RooPlot.h"
using namespace RooFit;
void fit2(double xl, double xh, int z=0)
{
// S e t u p m o d e l
// ---------------------
// Declare observable x
RooRealVar x("x", "x", 0, 10);
// Create two Gaussian PDFs g1(x,mean1,sigma) anf g2(x,mean2,sigma) and their parameters
RooRealVar mean("mean", "mean of gaussians", 5);
RooRealVar sigma1("sigma1", "width of gaussians", 0.5);
RooRealVar sigma2("sigma2", "width of gaussians", 1);
RooGaussian sig1("sig1", "Signal component 1", x, mean, sigma1);
RooGaussian sig2("sig2", "Signal component 2", x, mean, sigma2);
// Build Chebychev polynomial pdf
RooRealVar a0("a0", "a0", 0.5, 0., 1.);
RooRealVar a1("a1", "a1", 0.2, 0., 1.);
RooChebychev bkg("bkg", "Background", x, RooArgSet(a0, a1));
// Sum the signal components into a composite signal pdf
RooRealVar sig1frac("sig1frac", "fraction of component 1 in signal", 0.8, 0., 1.);
RooAddPdf sig("sig", "Signal", RooArgList(sig1, sig2), sig1frac);
// Sum the composite signal and background
RooRealVar bkgfrac("bkgfrac", "fraction of background", 0.5, 0., 1.);
RooAddPdf model("model", "g1+g2+a", RooArgList(bkg, sig), bkgfrac);
// C r e a t e b i n n e d d a t a s e t
// -----------------------------------------
std::unique_ptr<RooDataSet> d{model.generate(x, 10000)};
std::unique_ptr<RooDataHist> dh{d->binnedClone()};
// Construct a chi^2 of the data and the model.
// When a pdf is used in a chi^2 fit, the probability density scaled
// by the number of events in the dataset to obtain the fit function
// If model is an extended pdf, the expected number events is used
// instead of the observed number of events.
x.setRange("fitRange", xl, xh);
model.chi2FitTo(*dh, {PrintLevel(-1)}, Range("fitRange"));
// NB: It is also possible to fit a RooAbsReal function to a RooDataHist
// using chi2FitTo().
// Note that entries with zero bins are _not_ allowed
// for a proper chi^2 calculation and will give error
// messages
std::unique_ptr<RooAbsData> dsmall{d->reduce(EventRange(1, 100))};
std::unique_ptr<RooDataHist> dhsmall{static_cast<RooDataSet&>(*dsmall).binnedClone()};
std::unique_ptr<RooAbsReal> chi2_lowstat{model.createChi2(*dhsmall)};
ofstream myout;
myout.open("chi2_results.txt",ios::app);
myout << Form("sPlot_%.0f-%.0f_%d / ",xl,xh,z) << chi2_lowstat->getVal() << endl;
myout.close();
TCanvas *cdata = new TCanvas("sPlot", "sPlot demo", 400, 600);
RooPlot *frame = x.frame(Title("Fit of model to discriminating variable"));
dh->plotOn(frame);
if (z==0) model.plotOn(frame, Name("FullModel"));
else model.plotOn(frame, Name("FullModel"), Range("fitRange"));
frame->Draw();
cdata->SaveAs(Form("sPlot_%.0f-%.0f_%d.png",xl,xh,z));
}
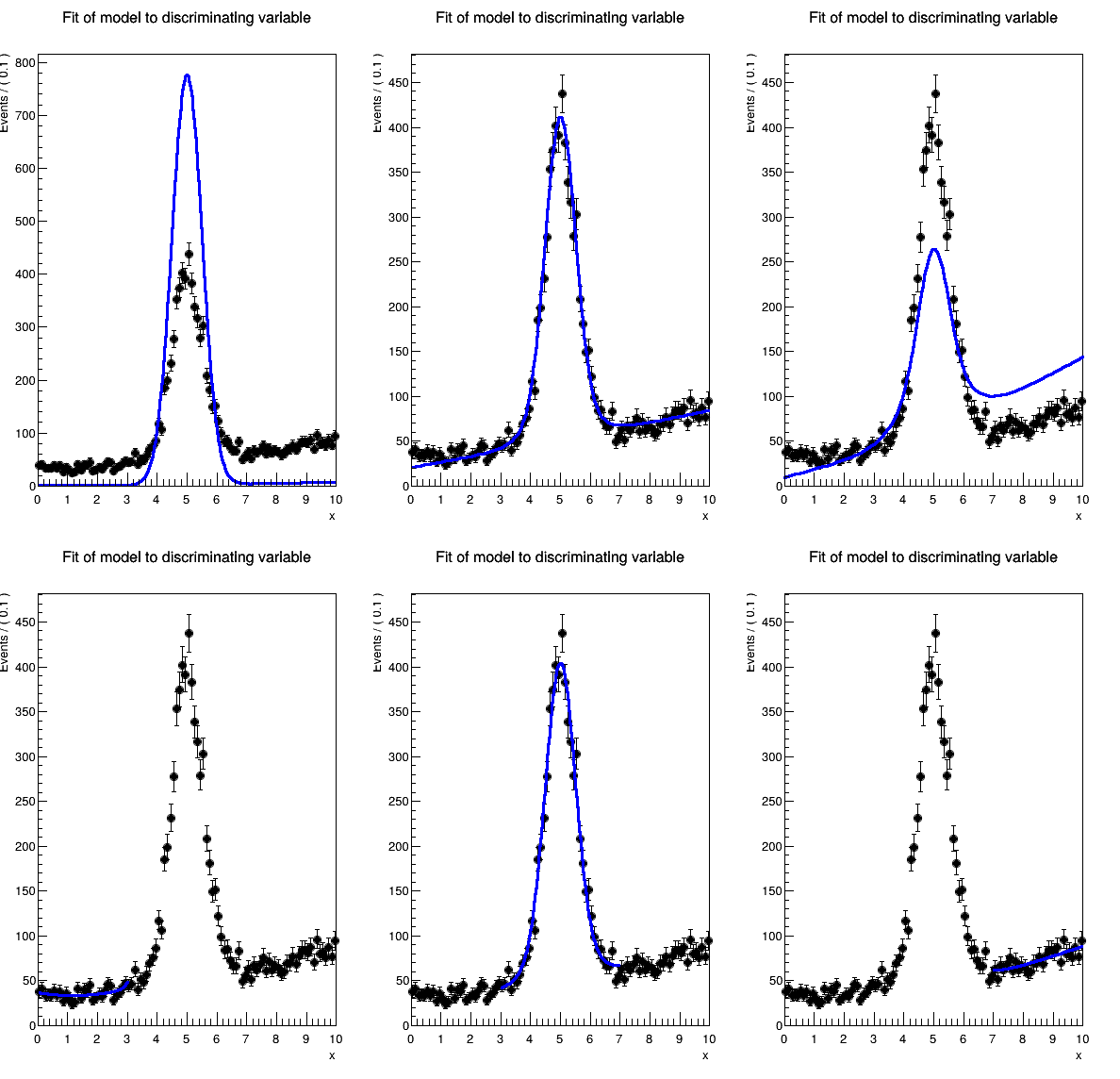
The results are:
where the top row is from plotOn without range (z=0 in the macro) and the bottom is with Range. The 3 columns are for the 3 ranges (0,3), (3,7), and (7,10). The results file contains:
sPlot_0-3_0 / 2385.35
sPlot_3-7_0 / 104.764
sPlot_7-10_0 / 145.656
sPlot_0-3_1 / 2385.35
sPlot_3-7_1 / 104.764
sPlot_7-10_1 / 145.656
So I think the fit is indeed using the range (but apparently not when using number limits directly in chi2FitTo, as you found, as the results don’t change with that), but the plot shows the whole model unless you add Range too in plotOn, and also note that plotOn normalises the model when plotting it (ROOT: RooAbsPdf Class Reference), so it can look different from what you would expect depending on what is done. Anyway, Jonas or someone else may provide more insights.
I added
model.getParameters(x)->Print("s") ;
to your code to see what the fit parameters look like and I ended up getting the same fit parameters for each ranged fit:
Processing fit2.C(0,3,0)...
[#0] WARNING:InputArguments -- The parameter 'sigma1' with range [-inf, inf] of the RooGaussian 'sig1' exceeds the safe range of (0, inf). Advise to limit its range.
[#0] WARNING:InputArguments -- The parameter 'sigma2' with range [-inf, inf] of the RooGaussian 'sig2' exceeds the safe range of (0, inf). Advise to limit its range.
[#1] INFO:Eval -- RooRealVar::setRange(x) new range named 'fitRange' created with bounds [0,3]
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: activating const optimization
[#1] INFO:Minimization -- The following expressions have been identified as constant and will be precalculated and cached: (sig1,sig2)
[#1] INFO:Minimization -- The following expressions will be evaluated in cache-and-track mode: (bkg)
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: deactivating const optimization
1) RooRealVar:: a0 = 0.501526 +/- 0.0229093
2) RooRealVar:: a1 = 0.158456 +/- 0.0368273
3) RooRealVar:: bkgfrac = 0.506609 +/- 0.0113469
4) RooRealVar:: mean = 5
5) RooRealVar:: sig1frac = 0.815448 +/- 0.0373635
6) RooRealVar:: sigma1 = 0.5
7) RooRealVar:: sigma2 = 1
Info in <TCanvas::Print>: png file sPlot_0-3_0.png has been created
Processing fit2.C(3,7,0)...
[#0] WARNING:InputArguments -- The parameter 'sigma1' with range [-inf, inf] of the RooGaussian 'sig1' exceeds the safe range of (0, inf). Advise to limit its range.
[#0] WARNING:InputArguments -- The parameter 'sigma2' with range [-inf, inf] of the RooGaussian 'sig2' exceeds the safe range of (0, inf). Advise to limit its range.
[#1] INFO:Eval -- RooRealVar::setRange(x) new range named 'fitRange' created with bounds [3,7]
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: activating const optimization
[#1] INFO:Minimization -- The following expressions have been identified as constant and will be precalculated and cached: (sig1,sig2)
[#1] INFO:Minimization -- The following expressions will be evaluated in cache-and-track mode: (bkg)
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: deactivating const optimization
1) RooRealVar:: a0 = 0.501526 +/- 0.0229093
2) RooRealVar:: a1 = 0.158456 +/- 0.0368273
3) RooRealVar:: bkgfrac = 0.506609 +/- 0.0113469
4) RooRealVar:: mean = 5
5) RooRealVar:: sig1frac = 0.815448 +/- 0.0373635
6) RooRealVar:: sigma1 = 0.5
7) RooRealVar:: sigma2 = 1
Info in <TCanvas::Print>: png file sPlot_3-7_0.png has been created
Processing fit2.C(7,10,0)...
[#0] WARNING:InputArguments -- The parameter 'sigma1' with range [-inf, inf] of the RooGaussian 'sig1' exceeds the safe range of (0, inf). Advise to limit its range.
[#0] WARNING:InputArguments -- The parameter 'sigma2' with range [-inf, inf] of the RooGaussian 'sig2' exceeds the safe range of (0, inf). Advise to limit its range.
[#1] INFO:Eval -- RooRealVar::setRange(x) new range named 'fitRange' created with bounds [7,10]
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: activating const optimization
[#1] INFO:Minimization -- The following expressions have been identified as constant and will be precalculated and cached: (sig1,sig2)
[#1] INFO:Minimization -- The following expressions will be evaluated in cache-and-track mode: (bkg)
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: deactivating const optimization
1) RooRealVar:: a0 = 0.501526 +/- 0.0229093
2) RooRealVar:: a1 = 0.158456 +/- 0.0368273
3) RooRealVar:: bkgfrac = 0.506609 +/- 0.0113469
4) RooRealVar:: mean = 5
5) RooRealVar:: sig1frac = 0.815448 +/- 0.0373635
6) RooRealVar:: sigma1 = 0.5
7) RooRealVar:: sigma2 = 1
Info in <TCanvas::Print>: png file sPlot_7-10_0.png has been created
Processing fit2.C(0,3,1)...
[#0] WARNING:InputArguments -- The parameter 'sigma1' with range [-inf, inf] of the RooGaussian 'sig1' exceeds the safe range of (0, inf). Advise to limit its range.
[#0] WARNING:InputArguments -- The parameter 'sigma2' with range [-inf, inf] of the RooGaussian 'sig2' exceeds the safe range of (0, inf). Advise to limit its range.
[#1] INFO:Eval -- RooRealVar::setRange(x) new range named 'fitRange' created with bounds [0,3]
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: activating const optimization
[#1] INFO:Minimization -- The following expressions have been identified as constant and will be precalculated and cached: (sig1,sig2)
[#1] INFO:Minimization -- The following expressions will be evaluated in cache-and-track mode: (bkg)
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: deactivating const optimization
1) RooRealVar:: a0 = 0.501526 +/- 0.0229093
2) RooRealVar:: a1 = 0.158456 +/- 0.0368273
3) RooRealVar:: bkgfrac = 0.506609 +/- 0.0113469
4) RooRealVar:: mean = 5
5) RooRealVar:: sig1frac = 0.815448 +/- 0.0373635
6) RooRealVar:: sigma1 = 0.5
7) RooRealVar:: sigma2 = 1
[#1] INFO:Plotting -- RooAbsPdf::plotOn(model) only plotting range 'fitRange', curve is normalized to data in given range
Info in <TCanvas::Print>: png file sPlot_0-3_1.png has been created
Processing fit2.C(3,7,1)...
[#0] WARNING:InputArguments -- The parameter 'sigma1' with range [-inf, inf] of the RooGaussian 'sig1' exceeds the safe range of (0, inf). Advise to limit its range.
[#0] WARNING:InputArguments -- The parameter 'sigma2' with range [-inf, inf] of the RooGaussian 'sig2' exceeds the safe range of (0, inf). Advise to limit its range.
[#1] INFO:Eval -- RooRealVar::setRange(x) new range named 'fitRange' created with bounds [3,7]
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: activating const optimization
[#1] INFO:Minimization -- The following expressions have been identified as constant and will be precalculated and cached: (sig1,sig2)
[#1] INFO:Minimization -- The following expressions will be evaluated in cache-and-track mode: (bkg)
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: deactivating const optimization
1) RooRealVar:: a0 = 0.501526 +/- 0.0229093
2) RooRealVar:: a1 = 0.158456 +/- 0.0368273
3) RooRealVar:: bkgfrac = 0.506609 +/- 0.0113469
4) RooRealVar:: mean = 5
5) RooRealVar:: sig1frac = 0.815448 +/- 0.0373635
6) RooRealVar:: sigma1 = 0.5
7) RooRealVar:: sigma2 = 1
[#1] INFO:Plotting -- RooAbsPdf::plotOn(model) only plotting range 'fitRange', curve is normalized to data in given range
Info in <TCanvas::Print>: png file sPlot_3-7_1.png has been created
Processing fit2.C(7,10,1)...
[#0] WARNING:InputArguments -- The parameter 'sigma1' with range [-inf, inf] of the RooGaussian 'sig1' exceeds the safe range of (0, inf). Advise to limit its range.
[#0] WARNING:InputArguments -- The parameter 'sigma2' with range [-inf, inf] of the RooGaussian 'sig2' exceeds the safe range of (0, inf). Advise to limit its range.
[#1] INFO:Eval -- RooRealVar::setRange(x) new range named 'fitRange' created with bounds [7,10]
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: activating const optimization
[#1] INFO:Minimization -- The following expressions have been identified as constant and will be precalculated and cached: (sig1,sig2)
[#1] INFO:Minimization -- The following expressions will be evaluated in cache-and-track mode: (bkg)
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: deactivating const optimization
1) RooRealVar:: a0 = 0.501526 +/- 0.0229093
2) RooRealVar:: a1 = 0.158456 +/- 0.0368273
3) RooRealVar:: bkgfrac = 0.506609 +/- 0.0113469
4) RooRealVar:: mean = 5
5) RooRealVar:: sig1frac = 0.815448 +/- 0.0373635
6) RooRealVar:: sigma1 = 0.5
7) RooRealVar:: sigma2 = 1
[#1] INFO:Plotting -- RooAbsPdf::plotOn(model) only plotting range 'fitRange', curve is normalized to data in given range
Info in <TCanvas::Print>: png file sPlot_7-10_1.png has been created
Which I don’t believe I should get if I am applying a range to the fit? I also get the same if I try to fit background alone, which definitely shouldn’t happen because the background would fail to fit in the middle region. I don’t think chi2FitTo is applying Range() to be fit within, even if added to plotOn
Hi, thanks for the post!
Indeed, the range argument has been completely ignored by the Chi2 fit routines in RooFit. But you were not the only one noticing this, and I have brought chi2 fitting to the same level of command argument feature support as the NLL fitting in the upcoming ROOT 6.40.
If you want to try it out, you can check out the ROOT nightlies:
Note that the 6.40 release candidate doesn’t have all the chi2 fit fixes in yet, since I made some improvements after the the release candidate was released. The final ROOT 6.40.00 will be released in mid May.
Cheers,
Jonas