Dear experts,
I am trying to use RooStats to set expected limit on the signal number of events.
I attached a folder with all the code I am using. There you can find:
-
the analysis.root file, where the signal and background (bkg_model_full__y) histograms are saved.
-
the do_workspace_pdf.C takes the signal and background (bkg_model_full__y) histograms from the analysis.root and build the workspace by adding them as pdf, with
w->import(N_data);
w->import(n_signal);
w->import(signal_pdf);
w->import(Bkg_Model_Full_pdf);
w->factory("SUM::total_model(N_data*Bkg_Model_Full_pdf,n_signal*signal_pdf)");
The output is the model_config_pdf.root file, that is the input to the limit_calc_pdf.C
The parameter of interest is the number of signal events, so after running the limit I find 1170, that is ~650 times the expected number of events (1.8)
- the do_workspace_histfunc.C takes the signal and background (bkg_model_full__y) histograms from the analysis.root and build the workspace by adding them as histfunc:
w->factory("binw[1.]") ;
w->factory("expr::S('mu*binw',mu[1.],binw)") ;
w->factory("expr::B_cr('B1*binw',B1[1.],binw)") ;
w->factory("ASUM::total_model(S*signal_hf,B_cr*Bkg_Model_Full_hf)") ;
The output is the model_config_hist_func.root file, that is the input to the limit_calc_histfunc.C
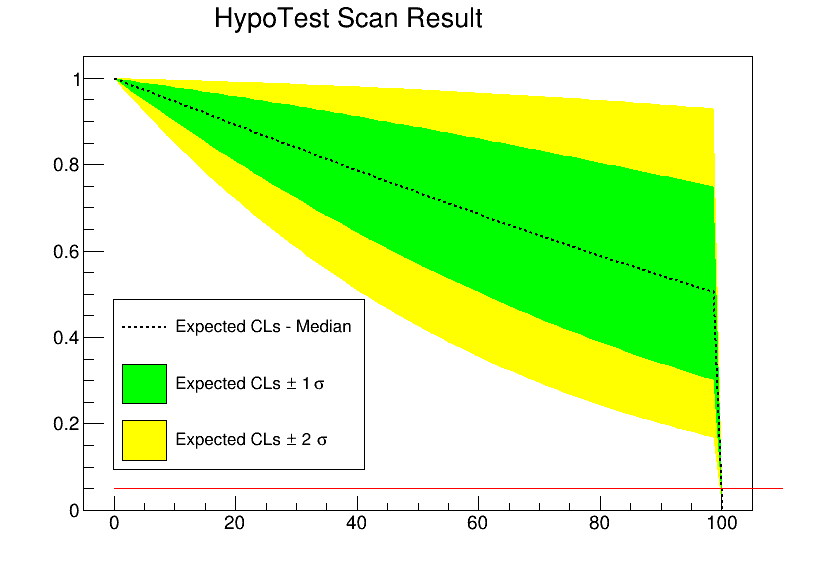
In this case parameter of interest should be the signal strength mu, but I am not able to run the limit, if I set binw=1 (I attached the plot here
I think that I should not change the binw, since I am always keeping the same number of bins and range.
However, I see that if I change the binw to 0.2, the CLs scan can be done, and an upper limit of 650 is obtained (that should be correct since here I am calculating upper limit on mu).
The only explanation I have is that my bin width is 5 GeV, so if I divide the number of events by 5, I am multiplying by 0.2… but I am not sure. I would like to know your opinion, to be sure that I am doing things correctly.
Thank you,
Laura
P.S. In the analysis.root file you see three background templates in it (bkg_model_full__y, HighHistForSyst4__y, LowHistForSyst4__y ) since I used a polynomial to describe the background, and they have been obtained by varying its coefficients of 0 (bkg_model_full__y) and +/- 1 sigma (HighHistForSyst4__y, LowHistForSyst4__y). The idea will be to use them to calculate systematics with PiecewiseIntepolation. However, on this point I would like to discuss later.
model_config_hist_func.root (14.5 KB)
limit_calc_histfunc.C (2.7 KB)
do_workspace_histfunc.C (3.7 KB)
do_workspace_pdf.C (2.9 KB)
model_config_pdf.root (14.5 KB)
limit_calc_pdf.C (2.6 KB)
analysis.root (6.7 KB)
histo_maker.C (18.1 KB)