Hi,all.
I am fitting a TGraph with user-Defined Functions (TF1).
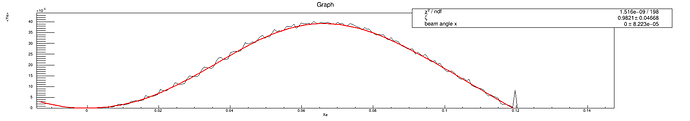
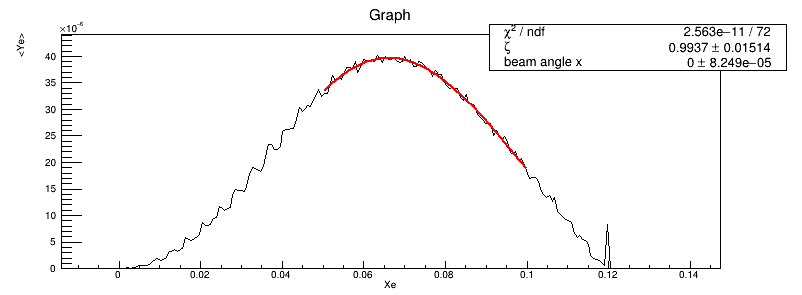
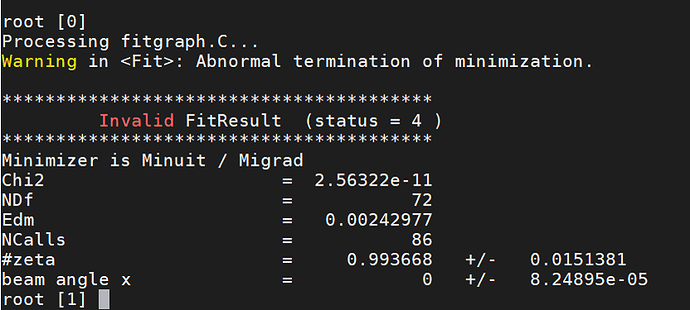
I am getting exceptionally large vales for chi2/NDF and I am confused, since the data fits the curve almost perfectly(see the plot).
The result is fit option "E"
The result is fit option"R"
I think the fit result is perfect from the figure. And the fit result shows the fit parameter(0.9937) is close to true value(equal to 1). But the only thing I don’t understand is that chi2/NDF is very large.
#include "TCanvas.h"
#include "TFrame.h"
#include "TBenchmark.h"
#include "TString.h"
#include "TF2.h"
#include "TH2.h"
#include "TFile.h"
#include "TROOT.h"
#include "TError.h"
#include "TInterpreter.h"
#include "TSystem.h"
#include "TPaveText.h"
//Define the fit fucntion Pai(y)
Double_t fitf(Double_t *x, Double_t *par)
{
double Ee = 45.5*1e9;
double w0 = 1.24;
double me = 0.511*1e6;
double kappa = 4*Ee*w0/(me*me);
double gamma = Ee/me;
double L1 = 60;
double L2 = 40;
double theta0 = 3.4467*1e-3;
double xx = (x[0]-par[1]*L1-kappa/2*theta0*L2)/(kappa/2*TMath::Sqrt(L1*L1/gamma/gamma+theta0*theta0*L2*L2));
double n = kappa/2;
double m = theta0*L2/TMath::Sqrt(L1*L1/gamma/gamma+theta0*theta0*L2*L2)*kappa/2;
double u = m*xx+n;
double AA = u*(1-xx*xx)*TMath::Pi();
double BB = 2*((1+(1+u)*(1+u)-4*u/kappa*(1+u)*(1-u/kappa))*TMath::Pi());
double value = par[0]*(L1/gamma*kappa/2)*(AA/BB);
return value;
}
void fitgraph() {
TCanvas *c1 = new TCanvas("c1_fit1","The Fit Canvas",200,10,700,500);
c1->SetGridx();
c1->SetGridy();
c1->GetFrame()->SetFillColor(21);
c1->GetFrame()->SetBorderMode(-1);
c1->GetFrame()->SetBorderSize(5);
gStyle->SetOptStat();
gStyle->SetOptFit();
// Pre-define histogram parameter
Double_t nbinsx = 200; // Bin number Y
Double_t nbinsy = 30; // Bin number Y
Double_t Xemin = -0.001;
Double_t Xemax = 0.135;
Double_t Yemin = -1*1e-3;
Double_t Yemax = 1*1e-3;
// Open the sample fiile
TFile *file1 = new TFile("Q_zheng10.root","read");
TFile *file2 = new TFile("Q_fu10.root","read");
TGraph *hzheng10, *hfu10;
hzheng10 = (TGraph *)file1->Get("Graph");
hfu10 = (TGraph *)file2->Get("Graph");
//Scale the histogram
double binwidth = (Xemax-Xemin)/200;
Double_t xaxis[200],yaxis[200];
for(Int_t n = 1; n<=200; n++)
{
xaxis[n] = Xemin + binwidth*(2*n-1)*0.5;
double *yzheng = hzheng10->GetY();
double *yfu = hfu10->GetY();
yaxis[n] = -0.5*yzheng[n] + 0.5*yfu[n];
}
c1->Divide(1,3);
c1->cd(1);
hzheng10->Draw();
c1->cd(2);
hfu10->Draw();
c1->cd(3);
TGraph *hcha_10 = new TGraph(nbinsx, xaxis, yaxis);
hcha_10->GetXaxis()->SetTitle("Xe");
hcha_10->GetYaxis()->SetTitle("<Ye>");
hcha_10->GetXaxis()->CenterTitle(true);
//3. Fit
TF1 *func = new TF1("func",fitf,0.05,0.1,2);
func->SetParNames("#zeta","beam angle x");
hcha_10->Draw();
hcha_10->Fit("func","R");
//func->Draw("same");
//func->SetLineColor(kRed);
}
Best regards,
Hurricane