Hi everyone,
I am trying to make the concolution of two functions. One is the exponential one (with two parameters that I want to leave free, to fit afterwards a histogram) multiplied by a function that looks like: f(x) = [0] exp(-[1]exp(-[2]*x)) + [3], whose parameters are obtained from a previous fit from another histogram (so each value contains value and error associated), the fnction resulting from the multiplication must be convoluted with a sum of two gaussians, each one again with parameters and errors, as they are obtained from a fit too.
I have been looking in tutorials and forums about doing convolution of such functions, but I could only find the convoution of functions in root (such as expo and gaus or landau), but in my case I need to consider all the errors associated to the other parameters and only leave free the two parameters of the exponential, and I don’t know how to fix the others. I’ve tried with FixParameter(), and also SetParError(), but even though the results (from the code I will write in a moment) does not make sense. I’m asking for help if someone knows how to do so, or if you have an idea of what I may be doing wrong!
TF1 *f1 = new TF1("f1","[0]*exp(-x/[1])*([2]*exp(-[3]*exp(-[4]*x)) + [5])");
f1.FixParameter(2,0.306010);
f1.SetParError(2,0.00671028);
//The same with the 3,4,5 parameters
TF1 *f2 = new TF1("f2","[6]*exp(-((x-[7])^2)/(2*([8]^2))) + [9]*exp(-((x-[7])^2)/(2*([10]^2)))");
f2.FixParameter(6,89.84);
f1.SetParError(6,21.50);
//The same with the 7,8,9,10 parameters
TF1Convolution *f_conv = new TF1Convolution("f1","f2",0.00,0.0194,true);
f_conv->SetRange(0.00,0.0194);
f_conv->SetNofPointsFFT(1000);
TF1 *ffit = new TF1("ffit",f_conv,0.00,0.0194,f_conv->GetNpar();
//Histogram we want to fit already uploaded by TFile, Get histogram, and named h
h->Sumw2();
//Trying to set an initial value for parameters of the exponential
ffit->SetParameter(0,10.);
ffit->SetParameter(1,620.);
h->Fit("ffit");
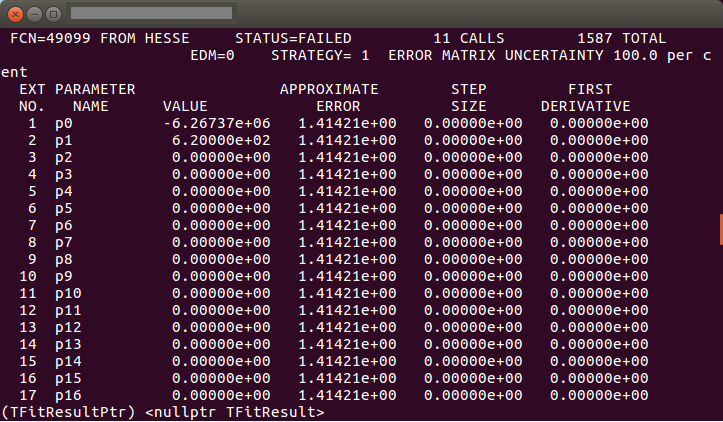
And the results I obtained are:
Also, I tried to put directly the values of the pre-fitter functions without considering the errors associated, so the functions f1 and f2 were something like: -234.234*x etc, what I mean is that I did not put the numbers as parameters, but writing directly the values of those. However, it didn’t work, and even though it returned only two parameters (both zero and with NaNs) it gave me the error Warning in <TCanvas::ResizePad>: Inf/NaN propagated to the pad. Check drawn objects. Warning in <TCanvas::ResizePad>: c1 height changed from 0 to 10 and I don’t know why can’t I make the concolution in any of those cases.
Thanks in advance!
L.