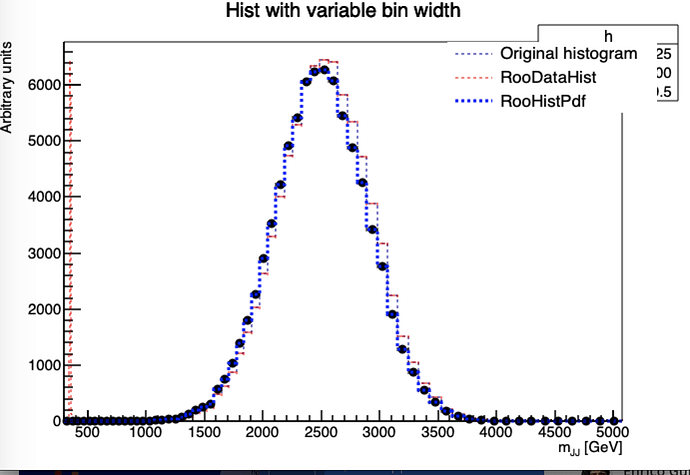
It’s a bit hacky, because you have to configure the plot options manually, but this is how you can do it:
32 RooAbsData::PlotOpt opts;
33 opts.bins = &mjjbin;
34 opts.correctForBinWidth = true;
35 opts.etype = RooAbsData::None;
36 hdata.plotOn(frameData, opts);
The corresponding function is here:
https://root.cern.ch/doc/master/classRooDataHist.html#acce2f981ef2374868e9c4d05f55aab12
and the PlotOpt here:
https://root.cern.ch/doc/master/structRooAbsData_1_1PlotOpt.html
Note:
This only works with the error types None and SumW2, since the default Poisson errors don’t work for non-integer bin counts. If the bin width correction is in effect, the bin counts are most likely not integers.