Hi,
There are two groups of TTrees that I would like to read together. For each TTree in the first group, there is a pair in the second group; however, the pair tree is not sorted and might have missing rows. So, I indexed rows using BuildIndex() then paired TTrees using Addfriend() and merged pairs by TChain().
I tried to give the chain to the RDataFrame but I noticed an unexpected behavior. When the parent TTree is the one with missing rows, the event loop looks fine. However, when the parent TTree is the bigger TTree, RDataFrame returns the leftover in the memory for the missing rows of the smaller TTree. Below is a demonstration for a single TTree pair.
# Define some numpy arrays:
run = np.arange(0,10, dtype = np.intc)
event = np.arange(0,10, dtype = np.intc)
rq = np.arange(100,110, dtype = np.intc)
rq_dict = {'run': run, 'event': event,'rq': rq}
rq_df = pd.DataFrame(rq_dict)
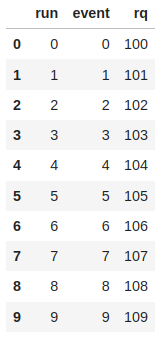
# A TTree from the first group
display(rq_df)
rrq_df = rq_df.sample(frac=0.5,random_state =1).drop('rq',axis = 1)
rrq_df['rrq'] = np.random.randint(1, 1000, rrq_df.shape[0])
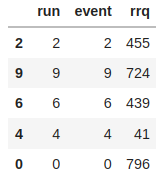
# A TTree from the second group(missing rows/shuffled)
display(rrq_df)
rrq_dict = {key:np.array(value,dtype = np.intc) for key, value in rrq_df.to_dict('list').items()}
# Makign ROOT files
rrq_df = ROOT.RDF.MakeNumpyDataFrame(rrq_dict)
rrq_df.Snapshot('zip1','rrq.root')
rq_df = ROOT.RDF.MakeNumpyDataFrame(rq_dict)
rq_df.Snapshot('zip1','rq.root')
rq_tree.BuildIndex('run','event')
rrq_tree.BuildIndex('run','event')
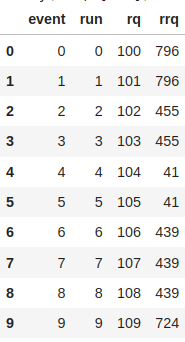
# Bigger TTree be the parent.
q_tree.AddFriend(rrq_tree)
df = ROOT.RDataFrame(rq_tree)
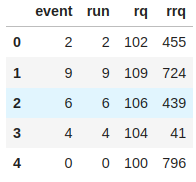
# Samller TTree be the parent.
rrq_tree.AddFriend(rq_tree)
df = ROOT.RDataFrame(rrq_tree)
Questions:
Is the behavior above expected?
I was hoping to get Nan for the missing rows. Is there a way to achieve that?
I would be happy to hear if there are other ideas on how to deal with such TTrees.
_ROOT Version: 6.26 - PyRoot