Hi,
I’m using
----- ROOT Version: 6.26/06
----- Python 3.8.13
I’m trying to merge two ROOT files with different branches (but same number of entries) in pyroot using AddFriend/dataframes/Snapshot. The file sizes are 50 MB and 200 kb. I’m using the following code:
f1 = ROOT.TFile.Open(“file_1.root”,“READ”) #50 MB
f2 = ROOT.TFile.Open(“file_2.root”,“READ”) #200 kB
t1=f1.Get(“nominal”)
t2=f2.Get(“nominal”)
t1.AddFriend(t2)
df = RDataFrame(t1)
df.Snapshot(“nominal”,“new_file.root”) #produces a file of size ~50MB, branches added etc, all good
f2.Close()
f1.Close()
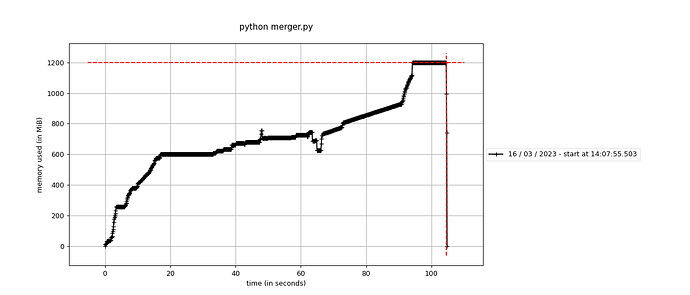
The code works as expected, but uses ~1.2 GB RAM, which is crazy! I mapped the RAM usage, in the first 0.5 seconds, everything up to the RDataframe creation is executed. The df.Snapshot takes ~100 second, and that’s the part where the memory blows
Any pointers on why this might be happening? Or a better workaround ![]()
While this is a test file, I’m dealing with files upto ~15 GB in size and such a blowup in RAM is not affordable with the system I use.
Cheers and thanks in advance!