Hello everyone,
actually I’m doing my first “large-Ttree” analysis in my phD and I need to use PyROOT because I will use some python packages to post-process the data. Actually I did a lot of analysis using C++ and I’m relatively new to PyROOT (but not at python).
The point is that, despite I expected lower speed with PyROOT than C++ compiled analysis program, I think that I can do a couple of things better to improve the speed over a Ttree loop but I’m not sure about how to do it.
First of all here you have the script im running: histos_for_calibrations.py (6.9 KB)
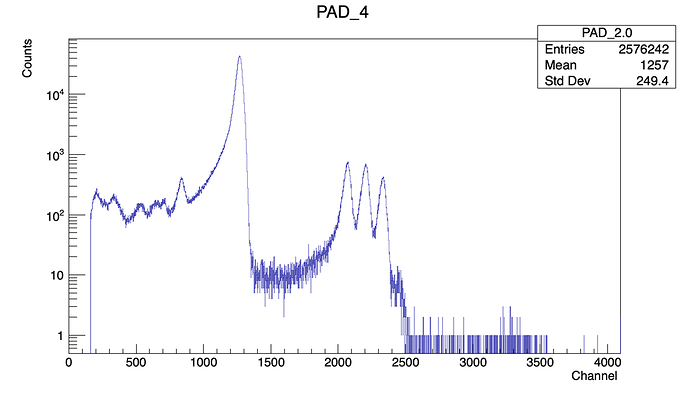
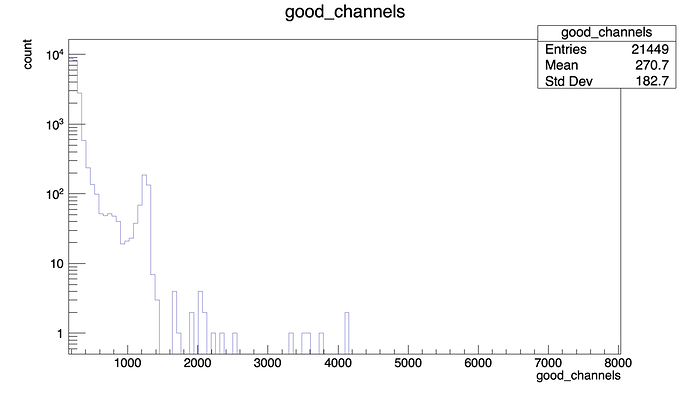
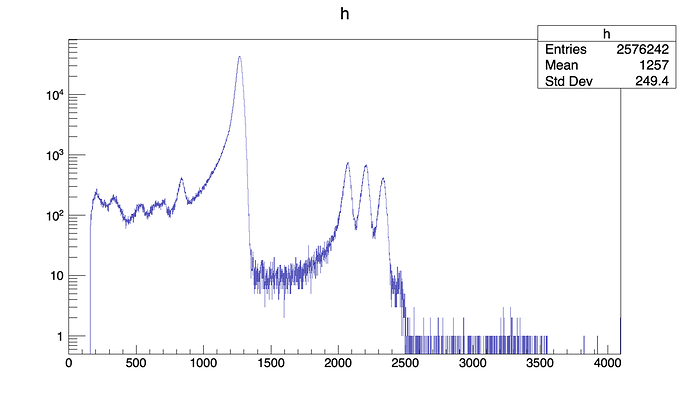
In the first part of the script I generate some empty histograms, then I fill them looping over the Ttree.
I read some tips in other posts of this forum but they are quite old and I couldn’t implement them to my code:
- First I tried was about SetBranchAddress of my Ttree as actually I access to my branches with something like:
tree.branch1. I feel that I will improve some speed if I declare previously each branch with SetBranchAddress but the point is that im not sure of how should I do it in python. For example with C++ I do:
UShort_t x[32];
tree->SetBranchAddress("x",x);
But If I try something similar in python like:
x = tree.SetBranchAddress('x', x)
It tells me that x is not declared previously. Do you have any idea of how to do it?
-
I read something about ROOT dataframe, which is used in PyROOT improve the speed quite a lot, but I couldn’t find any examples or how to use that dataframe. Could you help me with that?
-
The last point is that I also read something about compile python with pypy, but as I said, all info that I found was some years old and I’m not sure about that.
If there are some other methods different than the ones I mentioned above I will be glad to know it and try. Thank you very much for your help.