[quote=“dpiparo”]
2. Fit functions not displayed
I suspect you are fitting with the same function object the histograms. While mathematically this is perfectly sane, as you say the parameters are printed, in the program the function, and therefore the line on the canvas, is only one. Here you can find a proposal which uses 3 gaussians.
import ROOT
h1=ROOT.TH1F("h1","h1",64,-4,4)
h2=ROOT.TH1F("h2","h2",64,-4,4)
h3=ROOT.TH1F("h3","h3",64,-4,4)
# make them different
for h,s in zip([h1,h2,h3],[1,1.5,2]):
h.FillRandom("gaus")
h.Scale(s)
gfit1 = ROOT.TF1("gfit1","gaus(0)",-4,4)
gfit2 = ROOT.TF1("gfit2","gaus(0)",-4,4)
gfit3 = ROOT.TF1("gfit3","gaus(0)",-4,4)
h1.Fit(gfit1,"rme")
h2.Fit(gfit2,"rme")
h3.Fit(gfit3,"rme")
c=ROOT.TCanvas()
h3.Draw()
h2.Draw("Same")
h1.Draw("Same")
c.Draw()
Good luck with your internship 
Cheers,
D[/quote]
Hi there, thanks for your quick response. I have tried your suggestions and here are my results:
[ul]1. Your legend solution worked mostly - thanks! The only issue is that the colours of the plots do not show up on the legend. I have tried fixing this with no results.[/ul]
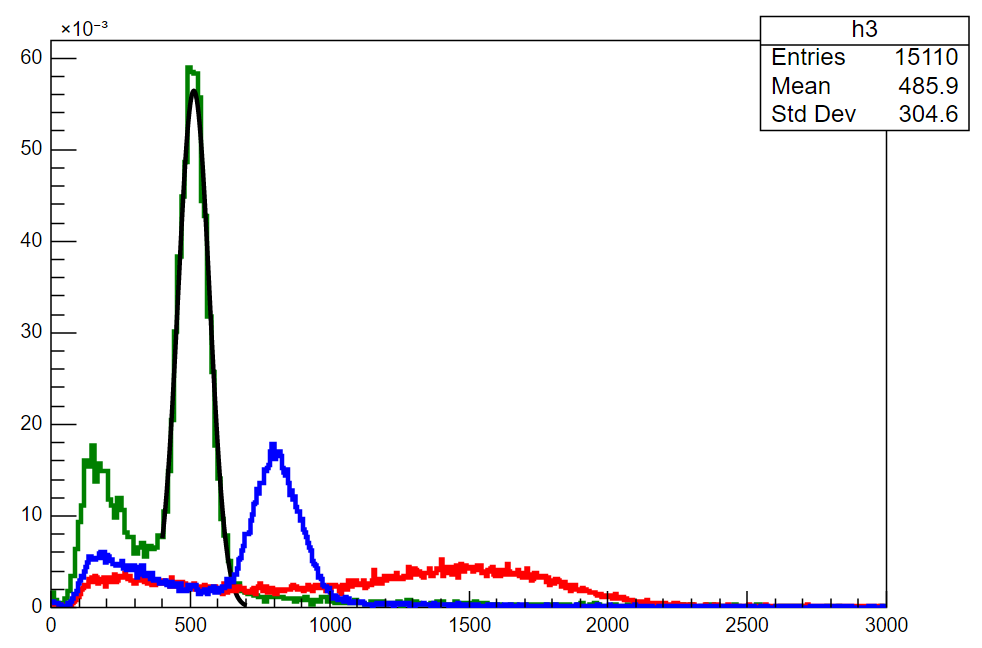
[ul]2. Your Gaussian solution seems to not have worked. I have provided a full copy of the code written by my colleague that gives the plot provided in my first post - as far as I can see, it is identical to the solution you provided sans the for-loop. I ran the code you provided, and like ours, it only plots the last-called Gaussian, gfit3. Could you elaborate on what exactly the for-loop is intending to do? It seems to arbitrarily scale them based on the tuple you make using zip(), which removes valuable information to us about the data. In our code below you can see that we have normalized our histograms so that we can compare them. [/ul]
[code]import ROOT
import pylab as pl
import numpy as np
ROOT.enableJSVis()
c1 = ROOT.TCanvas()
%matplotlib inline
Load our custom filters
ROOT.gSystem.Load("/usr/local/software/quadis/latest/quadis/build/lib/libquadis.so")
These are the names of our data files:
path1='qe12f000’
path2='qe12f001’
path3=‘qe12f002’
These are the locations of our data files to import and set pointers:
infile = ROOT.TFile("/data/news/queens/T1/"+path1+".root")
infile2 = ROOT.TFile("/data/news/queens/T1/"+path2+".root")
infile3 = ROOT.TFile("/data/news/queens/T1/"+path3+".root")
tree = infile.Get(“T1”)
tree2 = infile2.Get(“T1”)
tree3 = infile3.Get(“T1”)
tree.GetEntry()
tree2.GetEntry()
tree3.GetEntry()
nevent = ROOT.NEvent()
nevent2 = ROOT.NEvent()
nevent3 = ROOT.NEvent()
tree.SetBranchAddress(“NEvent”,nevent)
tree2.SetBranchAddress(“NEvent”,nevent2)
tree3.SetBranchAddress(“NEvent”,nevent3)
h1 = ROOT.TH1F(“h1”,“Normalized Amplitude Histogram”,1000,0,3000) #set range of h1
h2 = ROOT.TH1F(“h2”,“Normalized Amplitude Histogram”,1000,0,3000) #set range of h2
h3 = ROOT.TH1F(“h3”,“Normalized Amplitude Histogram”,1000,0,3000) #set range of h3
tree.Draw(“fPulseData[0].GetParameter(0)>>h1”)
tree2.Draw(“fPulseData[0].GetParameter(0)>>h2”)
tree3.Draw(“fPulseData[0].GetParameter(0)>>h3”)
gfit1 = ROOT.TF1(“gfit1”,“gaus(0)”,1000,1800)
gfit1.SetParameters(100,1500,1000,0,0)
gfit2 = ROOT.TF1(“gfit2”,“gaus(0)”,650,1000)
gfit2.SetParameters(100,750,500)
gfit3 = ROOT.TF1(“gfit3”,“gaus(0)”,400,700)
gfit3.SetParameters(100,500,500)
gfit1.SetLineColor(1)
gfit2.SetLineColor(1)
gfit3.SetLineColor(1)
Normalize all histograms
h1.Scale(1./h1.Integral())
h2.Scale(1./h2.Integral())
h3.Scale(1./h3.Integral())
h1.SetLineColor(2)
h2.SetLineColor(4)
h3.SetLineColor(3)
h1.Fit(gfit1,“rme”)
h2.Fit(gfit2,“rme”)
h3.Fit(gfit3,“rme”)
h3.Draw()
h1.Draw(“same”)
h2.Draw(“same”)
gfit1.Draw(“same”)
gfit2.Draw(“same”)
gfit3.Draw(“same”)
c1.Draw()
[/code]
Thanks again!